41
Diunggah oleh
ajquinonespHak Cipta
Format Tersedia
Bagikan dokumen Ini
Apakah menurut Anda dokumen ini bermanfaat?
Apakah konten ini tidak pantas?
Laporkan Dokumen IniHak Cipta:
Format Tersedia
41
Diunggah oleh
ajquinonespHak Cipta:
Format Tersedia
Connecting Medical Informatics and Bio-Informatics
R. Engelbrecht et al. (Eds.)
ENMI, 2005
971
Can openEHR Archetypes Empower
Multi-Centre Clinical Research?
Sebastian Gardea, Petra Knaupb, Thilo Schulera,c, Evelyn Hovengaa
a
Health Informatics Research Group, Central Queensland University, Rockhampton, Australia
b
University of Heidelberg, Department of Medical Informatics, Heidelberg, Germany
c
University of Freiburg, Department of Medical Informatics, Freiburg, Germany
Abstract
The Electronic Health Record is of utmost importance to enable the provision of
high-quality collaborative care; one prominent development is openEHR. On the
other hand, a systematic approach to support the use of routine data for multi-centre
clinical research is becoming increasingly important. One example of this is the
extensible architecture for using routine data for additional purposes (eardap) which
features comprehensive terminological support. However, as experiences in various
medical fields have shown, the terminology-based approach is limited to specialized
fields and it is argued that a comprehensive terminology is simply too complex and
too difficult to maintain. As the openEHR archetype approach does not rely heavily
on big standardized terminologies, it offers more flexibility during standardisation of
clinical concepts and overcome the shortcomings of terminology-focused approaches.
It is unknown, however, how far the more generic openEHR approach can also
enable re-use of routinely collected data for clinical research purposes the use case
for which eardap was designed. We therefore explored the feasibility of using the
openEHR approach to support multi-centre research in comparison to eardap.
Generally speaking, our results show that both eardap and openEHR are suitable to
enable the use of routine data for multi-centre clinical research. As the openEHR
approach also ensures open, future-proof Electronic Health Records, we conclude
that it is highly desirable that multi-centre clinical trials adopt openEHR.
Keywords:
Electronic Health Record, Terminology, openEHR, Medical Informatics, Clinical Trials
1. Introduction
The Electronic Health Record (EHR) is of utmost importance to provide high-quality
collaborative care. EHRs have the potential to offer simultaneous remote access to patient
data, increased legibility of the documents, flexible data layout and analysis, integration of
information resources, and tailored paper output [1]. In real life, however, a considerable
amount of information stored in the records is obsolete, redundant, duplicated, or
indecipherable or even contradictory to the extent that it does not benefit the patient at the
point of care. To solve this problem, several approaches are currently being explored. Two
prominent examples are the Clinical Document Architecture (CDA) [2] which is primarily
focussed on documents and document exchange, and the openEHR approach ([3],
http://www.openEHR.org) which focuses on the semantic interoperability of complete EHRs
or EHR extracts and is the basis for the new European standard [4].
Section 13: Public Health Informatics, Clinical Trials
Connecting Medical Informatics and Bio-Informatics
R. Engelbrecht et al. (Eds.)
ENMI, 2005
972
On the other hand, a systematic approach to support the use of routine data for clinical
research is becoming more important. To support clinical trials in medical fields where the
treatment is complex, the severity of the illness high or the incident rates low, collaborative
research efforts are vital and information technology support is essential. Such multi-centre
clinical trials may even cross national borders. There are powerful data collection,
management and Remote Data Entry (RDE) systems for clinical trials ([5], [6]) and even
multi-centre clinical trials ([7]). Some of them provide solutions for single terminological
aspects like the translation of Case Report Forms (CRFs), for the administration of
measurement units and conversion factors. However, there are very few offering a
comprehensive terminological support which is useful for conventional trials, desirable in
multi-centre trials and absolutely necessary in cooperative groups of multi-centre clinical
trials [8]. One example that supports comprehensive terminological support is eardap [9].
Still, the terminology-based approach is limited to specialized fields as research in various
medical fields has shown [10], [11] and it is argued that a comprehensive terminology is
simply too complex and too difficult to maintain [12]. As the openEHR archetype approach
does not rely on big standardized terminologies but micro-vocabularies [13], it would offer
more flexibility during standardisation of clinical concepts and overcome the shortcomings
of terminology-focused approaches. Further, openEHR provides the basis for future-proof,
medico-legally sound EHR systems. While this is not in the focus of eardap it might still be
valuable for ongoing multi-centre research.
It is unknown, however, how far the more generic openEHR approach for Electronic Health
Records can also enable the use of routine data for multi-centre research purposes the use
case eardap was designed for. We therefore explored the feasibility of the openEHR
approach to support this and compared its characteristics in detail with eardap.
The overall aim of this paper is to answer the research question - to what extent is openEHR
suitable for multi-centre research environments. We will
outline essential criteria for collaborative research environments,
show to what extent these criteria are fulfilled by eardap and openEHR, and
highlight differences, advantages and disadvantages between eardap and openEHR.
2. Material and methods
2.1.
openEHR and archetypes
The aim of openEHR is to enable the development of open specifications and software for
EHR systems. openEHR is based on the results of the GEHR-Project of the European Union.
GEHR is an acronym for Good European Health Record respectively later Good Electronic
Health Record. Following GEHR several projects extended and refined its results (e.g. the
Synapses and SynEx projects). All these projects influenced the openEHR architecture.
openEHR has pioneered a two level modelling approach for EHRs ([3]). An overview of this
approach is given in Figure 1. The first level is the reference information model which is
pared down to the minimum to support the medico-legal requirements and record
management functions. This ensures that clinicians can always send information to another
provider and receive information which they can read thus ensuring data interoperability.
The second level involves the openEHR archetype methodology a way of sharing evolving
clinical information so that it can be processed by the receiving provider thus ensuring
semantic interoperability. A blood pressure archetype for example represents a description of
all the information a clinician might want or has to report about a blood pressure
measurement. Basically, one archetype therefore represents one clinical concept.
Section 13: Public Health Informatics, Clinical Trials
Connecting Medical Informatics and Bio-Informatics
R. Engelbrecht et al. (Eds.)
ENMI, 2005
973
Figure 1: Overview of the openEHR two level modelling approach for EHRs ([3]).
Through the use of freely available archetype tools, e.g. the Archetype Editor, clinical
groups are empowered to control the way that EHRs are built up, using designed structures to
express the required clinical data and assuring that all necessary constraints on the values of
record components are observed. This ensures that all data in an EHR system is valid at two
levels, because it conforms both to an information model, and to domain-designed concept
definitions. Design principles of openEHR are described in more detail in [14], but the key
innovation of the openEHR architecture is that it separates record keeping concerns from
clinical data collection using archetypes [15] and thus enabling patient-centred,
longitudinal, comprehensive and prospective EHRs.
2.2.
eardap
eardap as an extensible architecture for using routine data for additional purposes was
developed to suit the needs of multi-centre clinical research in a multi-hospital environment
[9]. It focuses less on generic characteristics which Electronic Health Records must feature.
eardap can be characterized as terminology-based and component-based architecture.
eardap consists of 3 main components: core system, terminology management system
(TMS) and module generator (Figure 2). Main advantage of eardap is the comfortable
extensibility of any implemented architecture by new items and new research questions. Like
openEHR eardap is concept-oriented. In contrast, however, its architecture is based on
object-relational modelling supported by the TMS. The module generator is used each time a
new module has to be generated or an existing one has to be adapted. If the underlying
terminology has to be changed, the TMS is used: further modules will then be built upon the
changed terminology. Once the definition of a terminological system for a trial in the TMS is
finished, a consistent, corresponding relational database can be created within short time and
without any informatics skills. The process of building forms takes place under strict
terminological control. Generated research-specific modules can then be used by the eardap
core system in the medical centres.
3. Results
Based on our intensive requirements analyses with multi-centre trial environments (e.g. [8],
[16]), we developed criteria that are desirable for a multi-centre research environment. These
criteria are in harmony with other research (e.g. [5], [6], [17]) and are in the following
applied to both eardap and openEHR. For each criterion a description of how well it is
supported by either approach as well as an overall assessment is given (Table 1).
Assessments are given using the following scale: ++, +, +, , . As a baseline, + is given
Section 13: Public Health Informatics, Clinical Trials
Connecting Medical Informatics and Bio-Informatics
R. Engelbrecht et al. (Eds.)
ENMI, 2005
974
if the specification of the respective approach allows the criteria to be fulfilled but is either
not yet implemented or the scope of the implementation is outside the approach.
Figure 2: Overview of the eardap architecture in a typical eardap environment.
4. Discussion
Generally speaking, our results show that both eardap and openEHR are suitable to enable
the use of routine data for multi-centre medical research. eardap excels in providing highly
integrated tools for convenient analysis and report writing exclusively based on the
terminology provided and therefore usable for all scenarios. Further, mechanisms for high
data quality are supported by eardap through its warning and error integrity constraints.
openEHR excels in enabling a more flexible standardisation process and making
internationalisation and localisation feasible through concept-oriented multi-language
support and specialisation of archetypes. Further openEHR enables data and semantic
interoperability via its generic information model, archetypes and EHR extracts.
Our experiences in paediatric oncology in Germany have shown the applicability of the
eardap architecture for national research [9]. The functions of our core system including
additionally chemotherapy decision support based on the system were in routine use in
several hospitals all over Germany [16]. With eardap special emphasis has to be laid on
interfaces to local hospital information systems and data security.
The openEHR approach is currently being trialed in Australia in the framework of
HealthConnect (http://www.healthconnect.gov.au), the Australian initiative for a national
health information network and used in further projects [18]. First results are promising.
In this paper we have considered eardap and openEHR solely in the context of how well
they support clinical research based on routine clinical data; the primary purpose of
openEHR laying the foundation for sound Electronic Health Records - of course is slightly
different. Our criteria do not intend to generally assess approaches to Electronic Health
Records.
Independent of the approach used, the degree of reuse of routine data for multi-centre
research is highly dependant on the quality of the terminology used. Our experiences confirm
that terminology harmonization, maintenance and general governance are key factors for
Section 13: Public Health Informatics, Clinical Trials
Connecting Medical Informatics and Bio-Informatics
R. Engelbrecht et al. (Eds.)
ENMI, 2005
975
success in this area and a challenging task in a multi-centre project. The greater flexibility
during standardization processes offered by the openEHR archetyping is of great value here.
Table 1: Overview of all criteria and how well they are supported by eardap and openEHR.
Criteria
Usable for basic data
set documentation
Data
validation/integrity
constraints
eardap
Yes.
Sophisticated model for
warning and error integrity
constraints, intra- and
inter-contextual.
++
Support for multiple
trial terminologies1
Yes, inbuilt.
++
Support for evolving
terminologies
Yes, possible via new or
adapted research-specific
module based on terminology
server and created by eardap
module generator.
++
Automatic form
generation
Specialisation of
concepts allowed
Supports rapid
cross-patient analysis
Yes, via Form Building
Component.
Indirectly via specialized data
in research-specific module.
Integrated (standard and
flexible analysis based on
terminology).
Integrated (based on
terminology and templates).
Yes, but have to know
database schema.
++
Possible via HL7 or own
protocols. Context has to be
established.
Flexible through common
terminology (e.g. basic data
set) that is extendable by
research-specific
terminologies.
Only essential to agree on
basic data set by all parties.
Further items can be
standardised.
Not easily achievable.
Supports report writing
Provides data basis for
decision support
modules
Export/Import of Data
Degree of
standardization needed
Degree of governance
needed
Possibility for
internationalisation
(international trials)
++
+
++
++
+
openEHR
Yes.
Error integrity constraints can
easily be applied in one
archetype (one context). Warning
constraints or inter-contextual
constraints are indirectly
supported by templates and
invariants.
Not inherently supported by
openEHR. Achievable through
specialized archetypes and
external control which
archetypes are to be applied.
Yes, as openEHR features a
standard information model.
For incompatible changes a new
version of the archetype and
adequate update routines are
needed.
In the future, via templates,
GUI-Generator.
Yes (basic feature of archetypes).
Possible to implement even
retrospectively based on
archetypes, but not integrated.
Possible to implement, but not
integrated.
Yes, based on archetypes.
++
+
++
+
++
+
+
++
Possible via openEHR EHR
Extracts based on archetypes.
Context is guaranteed.
Even more flexible through
specialisation and because only
commonly used archetypes have
to be standardized.
++
Only essential to agree on
standardized archetypes that are
used by all parties, more flexible.
++
Yes, possible via context-based
translation of archetypes.
++
++
While HL7 primarily defines messages between applications and HL7 CDA is a generic
model for the communication of clinical documents, and in this is similar to openEHR
Transactions, openEHRs focus is the EHR as a whole. The Clinical Data Interchange
Standards Consortium Operational Data Model (CDISC ODM) as a format for clinical trial
1
Various trial terminologies extending the basic terminology and are applied based on patient characteristics
like diagnosis.
Section 13: Public Health Informatics, Clinical Trials
Connecting Medical Informatics and Bio-Informatics
R. Engelbrecht et al. (Eds.)
ENMI, 2005
976
data exchange could support data exchange between openEHR and non-openEHR clinical
trial systems.
5. Conclusion
It can be concluded that openEHR can support multi-centre research based on routine clinical
data about as well as eardap. As, in addition, openEHR inherently offers valuable features of
Electronic Health Records, we recommend that multi-centre clinical trials adopt the
openEHR approach for their research activities. For higher efficiency and data quality, some
of the features eardap excels in could be applied in addition to the openEHR methodology.
6. Acknowledgements
The authors wish to thank all those who have contributed with commitment and enthusiasm
to the openEHR and eardap projects.
7. References
[1] Powsner SM, Wyatt JC, and Wright P. Opportunities for and challenges of computerisation. Lancet 1998;
352(9140):1617-22.
[2] Dolin RH, Alschuler L, Beebe C, Biron PV, Boyer SL, Essin D, Kimber E, Lincoln T, and Mattison JE. The
HL7 Clinical Document Architecture. J Am Med Inform Assoc 2001; 8(6):552-69.
[3] Beale T. Archetypes: Constraint-based Domain Models for Future-proof Information Systems OOPSLA
2002 workshop on behavioural semantics., 2002.
[4] CEN/TC 251. EHRCOM prEN 13606-1. Health informatics Electronic health record communication
Part 1: Reference model. 2004.
[5] Duftschmid G, Gall W, Eigenbauer E, and Dorda W. Management of data from clinical trials using the
ArchiMed system. Med Inform Internet Med 2002; 27(2):85-98.
[6] Kuchinke W, Eich HP, and Ohmann C. Software for Remote Data Entry and Clinical Trials: Review of
Commercial Solutions and Trends 46. annual meeting of Deutsche Gesellschaft fr Medizinische
Informatik, Biometrie und Epidemiologie (gmds), Urban & Fischer, Cologne, 2001. 205-6.
[7] Abdellatif M and Reda DJ. A Paradox-based data collection and management system for multi-center
randomized clinical trials. Comput Methods Programs Biomed 2004; 73(2):145-64.
[8] Merzweiler A, Weber R, Garde S, Haux R, and Knaup-Gregori P. TERMTrial - Terminology-based
documentation systems for cooperative clinical trials. Comput Methods Programs Biomed 2005; to appear
[9] Knaup P, Garde S, Merzweiler A, Graf N, Weber R, and Haux R. Towards shared patient records: An
Architecture for Using Routine Data for Nationwide Research. EuroMISE 2004: EFMI Symposium on
Electronic Health Record, Health Registers and Telemedicine. Prague, 12.-15.4.2004. 2004;
[10] Brown PJ. Coming to terms with datasets for diabetes care. Diabetes Nutr Metab 2000; 13(4):215-9.
[11] Cimino JJ. Terminology tools: state of the art and practical lessons. Methods Inf Med 2001; 40(4):298-306.
[12] Rector AL. Terminology and concept representation languages: where are we? Artif Intell Med 1999;
15(1):1-4.
[13] Health Level 7. HL7 EHR System Functional Model: A Major Development Towards Consensus on
Electronic Health Record System Functionality - A White Paper. 2004.
[14] Beale T, Goodchild A, and Heard S. EHR Design Principles. openEHR Foundation; 2001.
[15] Goodchild A, Gibson K, Anderson L, and Bird L. The Brisbane Southside HealthConnect Trial:
Preliminary Results Health Informatics Conference (HIC), Brisbane, 2004.
[16] Garde S, Baumgarten B, Basu O, Graf N, Haux R, Herold R, Kutscha U, Schilling F, Selle B, Spiess C,
Wetter T, and Knaup P. A meta-model of chemotherapy planning in the
multi-hospital/multi-trial-center-environment of pediatric oncology. Methods Inf Med 2004;
43(2):171-83.
[17] Silva JS, Ball MJ, Chute CG, Douglas JV, Langlotz CP, C. NJ, and Scherlis WL. Cancer Informatics Essential Technologies for Clinical Trials. New York: Springer, 2002.
[18] Bird L, Goodchild A, and Tun Z. Experiences with a Two-Level Modelling Approach to Electronic Health
Records. Journal of Research and Practice in Information Technology 2003; 35(2)
Address for correspondence
Dr. Sebastian Garde, Health Informatics Research Group, Faculty of Informatics and Communication,
Central Queensland University, Rockhampton Qld 4702, Australia , s.garde@cqu.edu.au, Ph +61 (0)7 4930
6542, Fax +61 (0)7 4930 9729, http://infocom.cqu.edu.au/hi
Section 13: Public Health Informatics, Clinical Trials
Anda mungkin juga menyukai
- Introduction To Bonev Package: Dongmei Li February 12, 2016Dokumen3 halamanIntroduction To Bonev Package: Dongmei Li February 12, 2016ajquinonespBelum ada peringkat
- Bridgesampling: An R Package For Estimating Normalizing ConstantsDokumen31 halamanBridgesampling: An R Package For Estimating Normalizing ConstantsajquinonespBelum ada peringkat
- Bnstruct: An R Package For Bayesian Network Structure Learning With Missing DataDokumen26 halamanBnstruct: An R Package For Bayesian Network Structure Learning With Missing DataajquinonespBelum ada peringkat
- ARTP (Adaptive Rank Truncated Product) Package: Detailed Examples of Computing The Gene and Path-Way P-ValuesDokumen3 halamanARTP (Adaptive Rank Truncated Product) Package: Detailed Examples of Computing The Gene and Path-Way P-ValuesajquinonespBelum ada peringkat
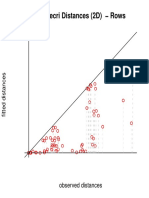
- Benzecri Distances (2D) RowsDokumen1 halamanBenzecri Distances (2D) RowsajquinonespBelum ada peringkat
- The Bmamevt Package: Bayesian Model Averaging at Work For Multivariate ExtremesDokumen5 halamanThe Bmamevt Package: Bayesian Model Averaging at Work For Multivariate ExtremesajquinonespBelum ada peringkat
- Jhs 11 FormsDokumen51 halamanJhs 11 FormsajquinonespBelum ada peringkat
- Spark 20 Tuning GuideDokumen21 halamanSpark 20 Tuning GuideajquinonespBelum ada peringkat
- Patient Demographics and MultivariableDokumen3 halamanPatient Demographics and MultivariableajquinonespBelum ada peringkat
- Debugging Tuning in SparkDokumen34 halamanDebugging Tuning in SparkajquinonespBelum ada peringkat
- Spark TuningDokumen26 halamanSpark TuningajquinonespBelum ada peringkat
- Data Mining of Agricultural Yield Data - A Comparison of Regression ModelsDokumen15 halamanData Mining of Agricultural Yield Data - A Comparison of Regression ModelsajquinonespBelum ada peringkat
- R PDF TablesDokumen4 halamanR PDF TablesajquinonespBelum ada peringkat
- R MapsDokumen36 halamanR MapsajquinonespBelum ada peringkat
- A Graphical User Interface Toolkit For RDokumen52 halamanA Graphical User Interface Toolkit For RajquinonespBelum ada peringkat
- TWacs BasicsDokumen116 halamanTWacs BasicsajquinonespBelum ada peringkat
- Shoe Dog: A Memoir by the Creator of NikeDari EverandShoe Dog: A Memoir by the Creator of NikePenilaian: 4.5 dari 5 bintang4.5/5 (537)
- The Yellow House: A Memoir (2019 National Book Award Winner)Dari EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Penilaian: 4 dari 5 bintang4/5 (98)
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeDari EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifePenilaian: 4 dari 5 bintang4/5 (5794)
- The Little Book of Hygge: Danish Secrets to Happy LivingDari EverandThe Little Book of Hygge: Danish Secrets to Happy LivingPenilaian: 3.5 dari 5 bintang3.5/5 (400)
- Grit: The Power of Passion and PerseveranceDari EverandGrit: The Power of Passion and PerseverancePenilaian: 4 dari 5 bintang4/5 (588)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureDari EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FuturePenilaian: 4.5 dari 5 bintang4.5/5 (474)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryDari EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryPenilaian: 3.5 dari 5 bintang3.5/5 (231)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceDari EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RacePenilaian: 4 dari 5 bintang4/5 (895)
- Team of Rivals: The Political Genius of Abraham LincolnDari EverandTeam of Rivals: The Political Genius of Abraham LincolnPenilaian: 4.5 dari 5 bintang4.5/5 (234)
- Never Split the Difference: Negotiating As If Your Life Depended On ItDari EverandNever Split the Difference: Negotiating As If Your Life Depended On ItPenilaian: 4.5 dari 5 bintang4.5/5 (838)
- The Emperor of All Maladies: A Biography of CancerDari EverandThe Emperor of All Maladies: A Biography of CancerPenilaian: 4.5 dari 5 bintang4.5/5 (271)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaDari EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaPenilaian: 4.5 dari 5 bintang4.5/5 (266)
- On Fire: The (Burning) Case for a Green New DealDari EverandOn Fire: The (Burning) Case for a Green New DealPenilaian: 4 dari 5 bintang4/5 (74)
- The Unwinding: An Inner History of the New AmericaDari EverandThe Unwinding: An Inner History of the New AmericaPenilaian: 4 dari 5 bintang4/5 (45)
- Rise of ISIS: A Threat We Can't IgnoreDari EverandRise of ISIS: A Threat We Can't IgnorePenilaian: 3.5 dari 5 bintang3.5/5 (137)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersDari EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersPenilaian: 4.5 dari 5 bintang4.5/5 (345)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyDari EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyPenilaian: 3.5 dari 5 bintang3.5/5 (2259)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreDari EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You ArePenilaian: 4 dari 5 bintang4/5 (1090)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)Dari EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Penilaian: 4.5 dari 5 bintang4.5/5 (121)
- Her Body and Other Parties: StoriesDari EverandHer Body and Other Parties: StoriesPenilaian: 4 dari 5 bintang4/5 (821)
- Azure Cosmos DB and DocumentDB SuccinctlyDokumen103 halamanAzure Cosmos DB and DocumentDB SuccinctlyRobert Kiefer100% (1)
- PBS BW AnalyserDokumen37 halamanPBS BW Analyserblaiso1Belum ada peringkat
- Batch Data Communication: BY - ArjunDokumen72 halamanBatch Data Communication: BY - ArjunKumar Ajit100% (1)
- Computer Mouse and Its TypesDokumen1 halamanComputer Mouse and Its TypesmbeaelnaaBelum ada peringkat
- Orange (Software)Dokumen6 halamanOrange (Software)levin696Belum ada peringkat
- Anti-Moussage GlycolDokumen6 halamanAnti-Moussage GlycolBn BnBelum ada peringkat
- Data Collection Scripts and Process Flow For Periodic Average Costing (PAC IPAC) - Cost ManagementDokumen13 halamanData Collection Scripts and Process Flow For Periodic Average Costing (PAC IPAC) - Cost ManagementLo JenniferBelum ada peringkat
- Manual de Taller Ford Ecosport 2006 170807031844Dokumen4 halamanManual de Taller Ford Ecosport 2006 170807031844Juan carlosBelum ada peringkat
- Modeling Data ObjectDokumen36 halamanModeling Data ObjectAsib KassayeBelum ada peringkat
- LUMIQ DataSheetDokumen5 halamanLUMIQ DataSheetBhanuj VermaBelum ada peringkat
- Zotero Vs MendeleyDokumen2 halamanZotero Vs Mendeleydzej_JJBelum ada peringkat
- SQLDokumen4 halamanSQLganiyamanBelum ada peringkat
- SIPI - Week 3 Review QuestionsDokumen12 halamanSIPI - Week 3 Review QuestionsAnggie OctaviaBelum ada peringkat
- Using TCDokumen770 halamanUsing TCkadher hussainBelum ada peringkat
- NetBackup 52xx and 53xx Appliance Admin Guide - 32Dokumen372 halamanNetBackup 52xx and 53xx Appliance Admin Guide - 32Avipan87Belum ada peringkat
- Maximo76 Cognos102 Install Guide Rev7Dokumen60 halamanMaximo76 Cognos102 Install Guide Rev7yuan_2425Belum ada peringkat
- DFo 6 1Dokumen17 halamanDFo 6 1Donna HernandezBelum ada peringkat
- New Dbms Lab ManualDokumen78 halamanNew Dbms Lab ManualDr. Hitesh MohapatraBelum ada peringkat
- Module 5 PDFDokumen68 halamanModule 5 PDFEum MavBelum ada peringkat
- Week Five Assignment Database Modeling and NormalizationDokumen9 halamanWeek Five Assignment Database Modeling and NormalizationEvans OduorBelum ada peringkat
- Yaqoob Et Al. - 2016 - Big Data From Beginning To FutureDokumen19 halamanYaqoob Et Al. - 2016 - Big Data From Beginning To FutureBruno OliveiraBelum ada peringkat
- List of Geographic Information Systems SoftwareDokumen8 halamanList of Geographic Information Systems SoftwareYankson Appiah DavidBelum ada peringkat
- DBMS With Mini Project 15CLS58 PDFDokumen70 halamanDBMS With Mini Project 15CLS58 PDFAnonymous jjuD0RFONBelum ada peringkat
- Final YearDokumen31 halamanFinal YearRAHULBelum ada peringkat
- 06a Memory v2Dokumen11 halaman06a Memory v2Vincent Terra NovaBelum ada peringkat
- Laboratory Activity No 1Dokumen4 halamanLaboratory Activity No 1Llal SantiagoBelum ada peringkat
- Softeng 351: Fundamentals of Database SystemsDokumen4 halamanSofteng 351: Fundamentals of Database SystemsAnonymous FUdMZrDyBelum ada peringkat
- Norton Chapter 01 AnswersDokumen5 halamanNorton Chapter 01 AnswersAmeena Ramzan79% (42)
- Etl Tools Informatica PDFDokumen2 halamanEtl Tools Informatica PDFBrettBelum ada peringkat
- Icon Design Lecture SlidesDokumen39 halamanIcon Design Lecture SlidesMinahel Noor FatimaBelum ada peringkat