Biol 130 Notes 2012
Diunggah oleh
Nick O'HaraHak Cipta
Format Tersedia
Bagikan dokumen Ini
Apakah menurut Anda dokumen ini bermanfaat?
Apakah konten ini tidak pantas?
Laporkan Dokumen IniHak Cipta:
Format Tersedia
Biol 130 Notes 2012
Diunggah oleh
Nick O'HaraHak Cipta:
Format Tersedia
B I O L O G Y 130
INTRODUCTORY CELL BIOLOGY
LECTURE NOTES
Course Author: Dr. N.C. Bols
Instructor: M. Pinheiro
Department of Biology
University of Waterloo
Fall, 2012
BIOL 130
LECTURE NOTES Fall, 2012
Lecture Notes
This booklet contains the notes that will be presented as part of the online modules. For
copyright reasons, the figures that will be shown along with the notes cannot be
reproduced. However, most of these figures come from the required course text, Cell and
Molecular Biology: Concepts and Experiments, 6th edition, Gerald Karp, John Wiley and
Sons, 2010. The notes point out where in the course text the figures that illustrate a
particular subject may be found. The exam questions come from the lecture notes.
Organization of Lecture Notes
There are 24 units or lectures (sometimes 1 unit takes less or more than the expected 1
lecture period). These 24 units or lectures are divided into six modules. Each module
begins with an outline for the module. A number, a letter and another number are used to
designate respectively a module, a lecture and a section. Thus 2f3 means module 2, unit f
of module 2 and section 3 of lecture f. These notations correspond to material on the
Learn website.
Using Lecture Notes
A useful way to use this booklet is to make notes in it and/or to make notes in it while
listening to the Biology 130 modules.
BIOL 130
LECTURE NOTES Fall, 2012
BIOLOGY 130 COURSE OUTLINE
INTRODUCTORY CELL BIOLOGY
Course Description: Introduction to the concepts of cell biology with an emphasis on
(1) the structural organization of the cell and (2) the function of critical molecular
processes that are characteristic of living organisms.
Course Objectives: At the end of the course you should be able to:
Explain some of the big concepts in cell biology, such as the difference between viruses
and cells, and the flow of genetic information.
Know in moderate detail several key pathways of energy metabolism and of signal
transduction.
Understand the vast vocabulary of cell biology so that the introductions will be easy to
many life science disciplines, such as biochemistry, molecular biology, genetics, and
physiology.
+++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
Related Course: BIOL 130L LAB 0.25 Cell Biology Laboratory. This course is run by
Dr. D. Miskovic (Ext. 35330) in ESC 357E (dmiskovi@sciborg.uwaterloo.ca). The
course could be taken concurrently with Biol 130, after Biol 130, or not at all.
+++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
ORDER OF MATERIAL TO BE COVERED AND EXAMINED ON
Modules (6) Subject (pages in Course Text )
Module 1: Introduction to Cells and Cellular Chemistry
1a Introduction to Cell Biology (1-30)
1b Chemical Basis of Life (31-39)
1c Lipids (39-41; 46-49)
1d Carbohydrates (42-46)
1e Nucleic acids (74-76; 386-390)
1f Proteins (49-63)
1
2
3
4
5
6
Module 2: Enzymes and Energy Metabolism
2a
2b
2c
Bioenergetics (84-92; 107; 182; 185-188)
Enzymes (92-103; 112-116)
Metabolism (105-116)
7
8
9
---------------------------------- Midterm on modules 1 & 2------------------------------------------------Module 3: Membranes and Energy Metabolism
3a Membranes (117-141; 250-258)
3b Membrane transport (143-155; 158-161)
3c Cellular uptake of particles & macromolecules (301-309)
3d ATP formation & the mitochondria (173-200)
3e Photosynthesis & the chloroplast (206-223)
10
11
12
13
14
BIOL 130
LECTURE NOTES Fall, 2012
Module 4: Flow of Genetic Information
4a Flow of information in cells (379-381; 386-390; 419-422; 455-457; 481-487)
4b Transcription & translation (422-455; 461-468)
4c Control of gene expression (503-531; 448-451)
4d DNA replication & repair (533-548; 552-554)
4e Cell cycle (560-571; 585-588; 590-591)
15
16
17
18
19
Module 5: Signal Transduction Pathways
5a Cell signaling & cAMP (605-614; 618-620)
5b Other 2nd messengers: lipids, calcium & nitric oxide (614-617; 634-638; 640-642)
5c Receptor Tyrosine Kinases, Cell Proliferation & Death (623-630; 638-640; 642-646)
20
21
22
Module 6: Biology of Cancer
6a Regulation of cell proliferation gone wrong (650-671)
6b Regulation of cellular social behavior gone wrong (230-258; 675-676)
23
24
---------------------------------Final exam will not cover Modules 1 & 2 -----------------------------------
BIOL 130
LECTURE NOTES Fall, 2012
PURCHASES TO MADE FROM UNIVERSITY BOOKSTORE FOR BIOLOGY 130:
Course text book Cell and Molecular Biology: concepts and experiments (2010 6th
edition) by G. Karp. The publisher is John Wiley & Sons Inc and the book is available
in the bookstore in two forms, listed below with ~ price.
i. Regular Bound Text (ISBN-13 978-0-470-48337-4).
~$149
The book will have resale value.
ii. Binder Ready version (ISBN 9780470556559) (Looseleaf)
~$97
This version might have little resale value.
The recommended readings from this text are listed in the course outline and in the
lecture notes.
used books
Earlier editions of the textbook can be used but they will have some disadvantages. The
pages in the course notes will not match precisely the pages in the older editions. The
older editions likely will have little subsequent resale value. The older editions will be
missing a few recent discoveries.
General Instructions about Tutorial Assignments
Tutorials for Biol 130 have been designed to help you develop and practice the skills that you
will need to use through your years of study at the University of Waterloo and not just in Biol
130. Some examples of these skills are:
to build up glossary and create concept maps
to efficiently use the Library
to read with understanding scientific journal articles
to write concisely and effectively
to orally present information on chosen relevant topics
BIOL 130
LECTURE NOTES Fall, 2012
University Policies to be aware of and sites to check
Academic Integrity: In order to maintain a culture of academic integrity, members of the
University of Waterloo community are expected to promote honesty, trust, fairness, respect
and responsibility. [Check www.uwaterloo.ca/academicintegrity/ for more information.]
Grievance: A student who believes that a decision affecting some aspect of his/her university
life has been unfair or unreasonable may have grounds for initiating a grievance. Read Policy
70, Student Petitions and Grievances, Section 4,
www.adm.uwaterloo.ca/infosec/Policies/policy70.htm. When in doubt please be certain to
contact the departments administrative assistant who will provide further assistance.
Discipline: A student is expected to know what constitutes academic integrity [check
www.uwaterloo.ca/academicintegrity/] to avoid committing an academic offence, and to take
responsibility for his/her actions. A student who is unsure whether an action constitutes an
offence, or who needs help in learning how to avoid offences (e.g., plagiarism, cheating) or
about rules for group work/collaboration should seek guidance from the course instructor,
academic advisor, or the undergraduate Associate Dean. For information on categories of
offences and types of penalties, students should refer to Policy 71, Student Discipline,
www.adm.uwaterloo.ca/infosec/Policies/policy71.htm. For typical penalties check Guidelines
for the Assessment of Penalties,
www.adm.uwaterloo.ca/infosec/guidelines/penaltyguidelines.htm.
Appeals: A decision made or penalty imposed under Policy 70 (Student Petitions and
Grievances) (other than a petition) or Policy 71 (Student Discipline) may be appealed if there
is a ground. A student who believes he/she has a ground for an appeal should refer to Policy 72
(Student Appeals) ww.adm.uwaterloo.ca/infosec/Policies/policy72.htm.
Note for Students with Disabilities: The Office for Persons with Disabilities (OPD), located
in Needles Hall, Room 1132, collaborates with all academic departments to arrange
appropriate accommodations for students with disabilities without compromising the academic
integrity of the curriculum. If you require academic accommodations to lessen the impact of
your disability, please register with the OPD at the beginning of each academic term.
BIOL 130
LECTURE NOTES Fall, 2012
Table of Contents for Biology 130 notes
Modules: Subject (Pages in Course Text)
Module 1: Introduction to Cells and Cellular Chemistry
1a Introduction to Cell Biology (1-30)
1b Chemical Basis of Life (31-39)
1c Lipids (38-41; 46-49)
1d Carbohydrates (42-46)
1e Nucleic acids (74-76; 386-390)
1f Proteins (49-63)
Module 2: Enzymes and energy metabolism
2a Bioenergetics (84-92; 107; 182; 185-188)
2b Enzymes (92-103; 112-116)
2c Metabolism (105-116)
Module 3: Membranes and energy metabolism
3a Membranes (117-141)
3b Membrane transport (143-155; 158-161)
3c Cellular uptake of macromolecules and particles (301-309)
3d ATP formation and the mitochondria (173-200)
3e Photosynthesis and the chloroplast (206-223)
Module 4: Flow of genetic information
4a Flow of information in cells (379-381; 386-390, 419-422; 455-457; 481-487)
4b Transcription and translation (422-455; 461-468)
4c Control of gene expression (503-531; 448-451)
4d DNA replication and repair (533-548; 552-554)
4e Cell cycle (560-571; 585-588; 590-591)
Module 5: Signal Transduction pathways
5a Cell signaling and cAMP (605-614; 618-6320)
5b Other 2nd messengers: lipids, calcium and nitric oxide (614-617; 634-638; 640-642)
5c Receptor Tyrosine Kinases, Cell Proliferation and Death (623-630; 638-640; 642-646)
Module 6: Biology of Cancer
6a Regulation of cell proliferation gone wrong (650-671)
6b Regulation of cellular social behavior gone wrong (230-258; 675-676)
BIOL 130
MODULE 1
Module 1 outline and notes: Introduction to Cells and Cellular Chemistry
1a Introduction to Cell Biology
1a1) Discovery and Properties of cells
1a2) Exceptions to the Cell Theory
1a3) Two fundamentally Different Classes of Cells
1a4) Eukaryotes vs. Prokaryotes
1a5) Common Features of Eukaryotes and Prokaryotes
1a6) Organisms from all species are made of cells
1a7) Types of Eukaryotic Cells
1b Chemical Basis of Life
1b1) Chemical Bonds
1b2) Polar Molecules
1b3) Ionization
1b4) Free Radicals
1b5) Biologically Important Weak Bonds
1b6) Nature of Biological Molecules
1b7) Water
1c Lipids
1c1) Introduction to four macromolecules
1c2) Lipids
1c3) Biological Roles of Lipids
1c4) Fatty acids
1c5) Triacylglycerols
1c6) Phosphoglycerides
1c7) Steroids
1d Carbohydrates
1d1) Introduction to Carbohydrates
1d2) Monosaccharides
1d3) Glucose and glucose
1d4) Disaccharides
1d5) Nutritional Polysaccharides
1d6) Structural Polysaccharides
1e Nucleic acids
1e1) Introduction to Nucleic Acids
1e2) Nitrogen Bases
1e3) Nucleosides
1e4) Nucleotides
1e5) Ribonucleic Acid (RNA)
1e6) Deoxyribonucleic Acid (DNA)
BIOL 130
MODULE 1
1f Proteins
1f1) Protein functions
1f2) Amino Acids
1f3) Peptide bond
1f4) Primary Structure of Proteins
1f5) Protein confirmation
1f6) Secondary Structure of Proteins
1f7) Tertiary Structure of Proteins
1f8) Motifs vs. Domains
1f9) Quaternary Structure of Proteins
1f10) Covalent Modifications of Proteins
1f11) Other Structural Features of Proteins
1f12) Multiprotein Complexes
BIOL 130
MODULE 1
1a: Introduction to Cell Biology: Unit #1
1a 1) Discovery and properties of cells (Karp: pp. 1-30)
History of cell theory
1.
2.
3.
4.
5.
Robert Hooke 1665
Leuwenhoek 1673-1700
Schleiden 1838-1839
Schwann 1838-1839
Rudolf Virchow 1858
Cell theory or cell doctrine
1.
2.
3.
All living things are composed of one or more units called cells.
Each cell is capable of maintaining its vitality independent of the rest
(i.e. Smallest clearly defined unit of life is the cell.)
Cells can arise only from other cells.
Basic Properties of Cells (living matter) (Karp: pp. 3-6)
1.
2.
3.
4.
5.
6.
7.
8.
Cells are highly complex and organized but all are enclosed by a physical barrier
cell membrane.
Blue print DNA (genetic program).
Cells acquire and utilize energy.
Cells carry out a variety of chemical reactions.
Cells are capable of producing more of themselves.
Cells engage in numerous mechanical activities.
Cells are able to respond to stimuli.
Cells are capable of self-regulation.
1a 2) Exceptions to the cell theory
Crossroads between living and non living matter
viruses (Karp: Figs. 1.20. 1.21, 1.22)
1. Viruses are bits of nucleic acids that have a protein coat
2. Inert outside the cells
3. Reproduce only in cells
viroids circular RNA without protein coat
prions proteinaceous infectious particles
BIOL 130
MODULE 1
1a 3) Two fundamentally different classes of cells
Prokaryotes before nucleus (Karp: p. 7 Fig. 1.8; p. 9 Table 1.1)
Always single cell organisms.
Prokaryotes 1-10 m (recently, rare exceptions have been found to be larger)
DNA lies free in cell or sometimes in an area called nucleoid region.
DNA is associated with fewer proteins and so is sometimes called 'naked'.
Eukaryotes true nucleus (Karp: p. 9 Table 1.1, p. 10 Fig. 1.10)
All cells of multicellular organisms are eukaryotes.
Eukaryotes 10-100 m
(exception, Caulerpa = largest single-cell organism)
DNA is organized into a nucleus.
Nucleus is an organelle.
DNA is associated with a characteristic set of proteins.
1a 4) Eukaryotes vs. Prokaryotes
Eukaryotes have but prokaryotes do not have the following:
1.
Organelles: Some examples
Mitochondria produce energy in form of ATP.
Lysosomes membrane bound sacs containing digestive enzymes.
Microbody (peroxisomes) oxidation of fatty acids and detoxification of
certain toxic compounds (hydrogen peroxide).
2.
Network of internal membranes:
Endoplasmic reticulum (e.r.)
rough e.r. ribosomes are bound to the membrane network.
smooth e.r. stores calcium
3.
Cytoskeleton
microtubules pipe-like cylinders about 20-25 nm in diameter.
microfilaments cylinders about 5 nm in diameter.
intermediate filaments cylinders or fibers 10 nm in diameter.
4.
Complex cilia and flagella
5.
Capacity for endocytosis and phagocytosis
1a 5) Common features of Eukaryotes and Prokaryotes (Karp: pp. 3-6)
BIOL 130
MODULE 1
1a 6) Organisms from all species are made of cells
Organisms can be divided into 6 kingdoms
(5 kingdoms if prokaryotes is considered just one).
Two prokaryote kingdoms (types of prokaryotic cells)
Archae (Archaebacteria)
often live in extreme environments.
e.g. thermophiles
all have cell walls.
Bacteria (Eubacteria) (Monera)
all have cell walls except mycoplasma.
mycoplasma are smallest cells (0.2 m)
Four eukaryote kingdoms
Protista (Figs. 1.16)
one celled and some colonial eukaryotic organisms.
Includes algae, water molds, slime molds and protozoa.
Fungi
Includes both multicellular organisms (e.g. mushrooms) and single-celled
organisms (e.g. yeast).
All have cell walls and are heterotrophs (depend on an external source of
organic compounds).
Plantae Always multicellular and always have cell walls
Most carry on photosynthesis.
Animalia
Always multicellular and hetertrophs (depend on an external source of organic
compounds).
1a 7) Types of eukaryotic cells
A.
Most complex single-celled (unicellular) protists (Fig. 1.16)
B.
Plant vs. Animal Cells (Karp: p. 8 Fig. 1.8)
Plants have
1. Cell walls made of cellulose and lignin
2. Plastids organelles bound by 2 membranes
concerned with energy metabolism e.g. chloroplast
3. Large vacuoles
C.
Cell specialization in multicellular organisms (differentiation) (Fig 1.17)
BIOL 130
MODULE 1
1b: Chemical Basis of Life: Unit #2
(Karp: p. 31-39)
1b 1) Chemical bonds
Hold two or more atoms together in an aggregate
Covalent bonds
The important strong bond in biological systems.
A chemical bond that results from sharing of electron pairs between atoms.
Covalent bonds can be formed between similar or even identical atoms.
An atomic aggregate linked together by covalent bonds is called a molecule.
Convention is to indicate a single covalent bond by a solid line between bonded
atoms. H-O-H
Common elements and their covalent bonding ability
1.
principal is that an atom is most stable when its outermost electron shell is filled.
2.
number of bonds depends on number of electrons needed to fill its outer shell.
(Karp: p. 32 Fig. 2.1)
C = carbon 4 covalent bonds or their equivalent in double and triple bonds
H = hydrogen 1 bond
O = oxygen 2 bonds
S = sulfur 2 bonds in organic molecules
N = nitrogen 3 bonds
P = phosphorus 5 (3) bonds
1b 2) Polar molecules
1.
2.
3.
4.
Certain atoms attract the shared electron pairs to a greater extent than other atoms.
This property of attracting electrons is called electronegativity.
When a covalent bond has an uneven distribution of charge, it is called a polar bond.
If molecule is appropriately shaped, a polar bond may result in a polar molecule or
dipole.
BIOL 130
MODULE 1
Polar and nonpolar molecules
Examples:
1.
2.
3.
H2O because of the angles of its bonds the H2O molecule is not a linear molecule
but, instead, has an angular shape.
This gives H2O molecule a partially negative end and two partially positive wings
so H2O is a polar molecule.
Molecules in which there is little or no separation of negative and positive charges
are nonpolar e.g. O2.
1b 3) Ionization
One atom loses electrons and another atom gains them.
Occurs because at least one partner is very electronegative.
Results in charged atoms or ions.
Negatively charged, anions.
(has extra electron relative to number of protons in nucleus)
Positively charged, cations.
(has extra proton relative to number of electrons)
Na+ and Cl- ions are stable because they possess filled outer shells
1b 4) Free radicals
Atoms or molecules that have orbitals containing a single unpaired electron.
Example is superoxide radical: O2.-
Highly reactive and damaging.
Might contribute to the process of aging.
Reactive oxygen species (ROS) are important free radicals.
1b 5) Biologically important weak bonds
(i.e. noncovalent bonds)
1.
2.
3.
Ionic bonds
Hydrogen bonds
Hydrophobic bond or hydrophobic interactions
Ionic bonds
Electrostatic attraction between fully charged components.
An example is table salt.
Attraction between positively charged Na+ and negatively charged Cl-.
BIOL 130
MODULE 1
Hydrogen bond (Karp: p. 36 & 37 Figs. 2.3, 2.7)
1.
2.
3.
4.
5.
A bond between an electronegative atom and a hydrogen atom that is already
covalently linked to another electronegative atom.
Hydrogen commonly bonds with highly electronegative atoms such as O and N.
Thus the pair of shared electrons is closer to the electronegative atom.
The positively charged nucleus of H atoms is readily attracted to unshared pair of
electrons of a second electronegative atom.
The noncovalent associations resulting from such electrostatic forces are called
hydrogen bonds.
Hydrophobic bonds (water-fearing) (Karp: p. 36 Fig. 2.5)
Tendency of nonpolar groups to aggregate when in the presence of H2O.
Nonpolar molecules are essentially insoluble in water because they lack the charged
regions that would attract them to the poles of water molecules.
In water, the nonpolar molecules are forced to aggregate.
1b 6) Nature of biological molecules
1.
2.
3.
Organic molecules: carbon-containing molecules.
Hydrocarbons: contain only carbon and hydrogen.
Most organic molecules in biology: are hydrocarbons with certain of the hydrogen
atoms replaced by various functional groups.
Functional groups:
1.
2.
3.
Particular groupings of atoms that often behave as a unit and give organic
molecules their properties.
Common functional groups (Karp: p. 41 Table 2.3 structural formula)
Examples: CH3, OH, COOH, NH2 (these are condensed structural form.)
Common linkage between functional groups.
Ester bonds formed between carboxylic acid and alcohols.
Amide bonds formed between carboxylic acid and amines.
1b 7) Water: Life-supporting properties of water
Fluid matrix around which insoluble fabric of cell is constructed.
Important because forms weak interactions with so many different chemical groups.
Water as a solvent
Dipolar nature of H2O makes it an ideal solvent for a variety of substances.
1.
Salts dissolve readily in H2O. e.g. NaCl (Karp: p. 35 Fig. 2.2)
2.
Covalent compounds with weakly polar properties
BIOL 130
3.
MODULE 1
Readily dissolved in H2O.
Compounds such as sugars have OH groups and carbonyl groups.
These dissolve because H2O molecules form hydrogen bonds with these polar
groups.
Nonpolar covalent molecules insoluble in water.
Such as benzene, ether, and chloroform are readily miscible only with one
another or with other nonpolar solvents.
BIOL 130
MODULE 1
10
1c: Lipids: Unit #3
(Karp: pp. 39-41; 46-49)
1c 1) Introduction to four macromolecules
Macromolecules huge, highly organized molecules that form the structure and carry
out the activities of cells.
Macromolecules can be divided into 4 major categories (Karp: p. 42 Fig. 2.11)
1. Lipids
2. Carbohydrates
3. Nucleic acids
4. Proteins
Carbohydrates, nucleic acids and proteins are polymers.
Polymers are composed of a large number of low-molecular-weight building
blocks, or monomers.
1c 2) Lipids
1.
2.
Small, diverse organic molecules that are insoluble in H2O but soluble in nonpolar
organic solvents. e.g. chloroform or benzene.
Hydrophobic or contain significant hydrophobic regions.
1c 3)_Biological roles of lipids
1.
2.
3.
4.
Source of energy in the diet and serve to store energy in the body.
e.g. fats and oils
Some hormones (chemical messengers) are lipids.
e.g. steroids and prostaglandins.
Prostaglandins usually act in an autocrine and paracrine fashion whereas steroid
hormones act in an endocrine fashion
Many vitamins are lipids.
e.g. vitamins A, D, E
The basic structural elements of biological membranes.
e.g. phospholipids
1c 4) Fatty acids (Karp: p. 47 Fig. 2.19b)
Unbranched hydrocarbon chains with a carboxyl group at one end.
Chains are typically 14 to 20 carbons.
Chain is hydrophobic.
Carboxyl group is hydrophilic.
Therefore fatty acids are amphipathic.
They can form micelles in water (Karp: p. 47 Fig. 2.20).
They may be saturated or unsaturated.
BIOL 130
MODULE 1
11
Saturated vs. unsaturated fatty acids
When all carbon atoms of a fatty acid chain are joined by single covalent bonds, the
compound is saturated.
O
CH3 - (CH2) n - C - OH
If one or more double bonds are present between carbons in the chain, the
compound is unsaturated.
O
CH3 - C = C - (CH2) n - C - OH
H H
1c 5) Fats and oils (triacylglycerols) (Karp: Fig. 2.19a, c & d)
Major compound for storing energy in both animals and plants.
Consist of glycerol esterified to three fatty acids.
The fatty acids can be identical.
If fatty acids are different, it is a mixed fat.
Fats that are liquid at room temperature are oils.
Oils contain unsaturated fatty acids.
1c 6) Phosphoglycerides (Karp: p 47 Fig. 2.22; p.123 Fig. 4.6)
Major component of membranes of all types.
Consist of glycerol esterified to two fatty acids.
The 3rd OH group of glycerol is bonded covalently to a phosphate group.
This is the parent compound, phosphatidic acid.
Several different, small polar groups can be linked to the phosphate.
ex. choline.
This would be phosphatidyl choline.
All phosphoglycerids are phospholipids.
All phospholipids are amphipathic.
1c 7) Steroids (Karp: p. 48 Fig. 2.21; p.123 Fig. 4.7)
Includes cholesterol which is found in membranes.
Cholesterol needed for synthesis of sex hormones.
Male androgens (e.g. testosterone) and female estrogens (estrogen)
BIOL 130
MODULE 1
1d: Carbohydrates: Unit #4
(Karp: pp. 42-46)
1d 1) Introduction to Carbohydrates
Have general formula (CH2O)n
Includes simple sugars (monosaccharides) and all larger molecules constructed of
sugar building blocks.
1.
Monosaccharides (simple sugars)
2.
Oligosaccharides 2 to 10 monosaccharide units linked together.
Attached to lipids: forms glycolipids.
Attached to proteins: forms glycoproteins
3.
Polysaccharides very long chains of monosaccharide units.
1d 2) Monosaccharides
Energy source and source of carbon for other cellular compounds.
Carbon chain containing one aldehyde or ketone
Plus (-OH) groups at the other carbons
aldehyde
ketone
HC=O
R1
C=0
R2
Aldehyde carbon is numbered #1.
Classified according to the number of C in the molecule.
Generally 3 to 7 carbon atoms.
Heptose has 7 carbons.
Hexose has 6 carbons.
Pentose has 5 carbons.
12
BIOL 130
MODULE 1
13
1d 3) - and -glucose: background
Stereoisomers
Isomers which have the same bonding sequence but which differ in how the atoms
are arranged in space.
Tetrahedral nature of carbon atom can lead to asymmetry in many organic
molecules (Karp: p. 43 Fig. 2.13).
This can lead to stereoisomers
Ring structures of monosaccharides
1.
2.
3.
Only very small amounts of monosaccharides are found in open chain form.
Most molecules are heterocyclic ring structures.
Rings result from intramolecular reactions between very active carbonyl group and
OH group on next to last carbon.
- and -glucose (Karp: p. 42 Fig. 2.12; p. 44 Fig. 2.15)
1.
2.
3.
4.
5.
Glucose is found as a six-membered ring (pyranose ring).
Ring formation introduces a new asymmetric carbon at position 1 (anomeric carbon),
so there is an additional pair of isomers.
These are stereoisomers that differ in configuration only at the anomeric carbon.
If the hydroxyl at position 1 is below the plane of the molecule, it is the anomer.
If the hydroxyl at position 1 is above the plane of the molecule, it is the anomer
1d 4) Disaccharides (Karp: p. 44 Fig. 2.16)
Consist of two monosaccharides linked together.
Covalent bond linking monosaccharides together is glycosidic bond.
e.g. Lactose = glucose and galactose
- l,4 linkage or glycosidic bond
e.g. Sucrose = glucose and fructose
- l,2 glycosidic bond
e.g. Maltose = 2 glucoses
- l,4 glycosidic bond
e.g. Cellobiose = 2 glucoses
- l,4 glycosidic bond
BIOL 130
MODULE 1
1d 5) Nutritional polysaccharides: glycogen and starch
Except at branch points, the repeating disaccharide is maltose.
Glycogen (Karp: p. 45 Fig. 2.17a)
Principal food reserve in animals and fungi: usually stored in liver and muscle of
animals.
-glucose units, mostly linked 1-4, but highly branched via frequent 1-6 linkages
Starches (Karp: p. 45 Fig. 2.17b)
Principal food reserve in plants.
Comes in 2 forms: amylose and amylopectin.
Amylose is an unbranched 1,4 polymer of glucose.
Amylopectin has same structure but is slightly branched.
Amylopectin is not as highly branched as glycogen.
1d 6) Structural polysaccharides
Cellulose (Karp: p. 45 Fig. 2.17c)
Linear polymer of several hundred to thousand glucose units.
An insoluble, rigid structural polymer.
Makes up cell wall of plants.
Cellobiose is the repeating unit of cellulose.
We cannot hydrolyze 1,4 linkage
Chitin (Fig. 2.18)
Unbranched polymer of the sugar N-acetylglucosamine.
N-acetylglucosamine like glucose except has acetylamino group at position 2.
Outer covering of insects, spiders and crustaceans.
Glycosaminoglycans (GAGs) (Karp, p. 46)
Made up of repeating dissacharide in which the two sugars are different.
One of the two sugar residues is always an amino sugar,
Either N-acetylglucosamine or N-acetylgalactosamine.
GAGs are found in extracellular matrix (ECM) of animal tissues.
14
BIOL 130
MODULE 1
15
1e: Nucleic Acids Unit #5
(Karp p. 74-76; 386-390)
1e 1) Introduction to Nucleic acids
Macromolecules constructed as a long chain (strand) of monomers, called nucleotides.
Either deoxyribonucleic acid (DNA) or ribonucleic acid (RNA).
Both are information molecules
Nucleotides (Karp: p. 75 Fig. 2.53a)
Consist of 3 units:
A nitrogen base
A pentose sugar
A phosphate group
1e 2) Nitrogen bases (Karp: p. 75 Fig. 2.54)
Organic compounds composed of C, N, H and in some cases O
There are two broad types:
Pyrimidine bases
three main bases
1. cytosine
2. uracil
3. thymine
Purine bases
two main bases
1. adenine
2. guanine
1e 3) Nucleosides
Pentose sugar 5 carbon sugar
Can be either ribose or deoxyribose
Purine or pyrmidine nucleosides
A purine or pyrimidine attached to a sugar.
IF the pentose is ribose, compound is a ribonucleoside. e.g. adenosine
IF pentose is deoxyribose, compound is deoxyribonuceloside. e.g. deoxyadenosine
BIOL 130
MODULE 1
16
1e 4) Nucleotides (Karp: p 75 Fig. 2.53a; p 386 Fig. 10.9b)
Nucleoside and phosphoric acid
Some examples:
Adenosine Monophosphate (AMP)
Adenosine Diphosphate (ADP)
Adenosine Triphosphate (ATP)
deoxyadenosine monophosphate (dAMP)
deoxyadenosine diphosphate (dADP)
deoxyadenosine diphosphate (dATP)
3', 5' cyclic adenosine monophosphate (cAMP)
Nucleotides are involved in three major cellular functions
1. Nucleotides are monomeric units from which DNA and RNA are made.
2. Second messengers in cell signaling e.g. cAMP.
3. They act as agents in energy-transferring reactions during metabolism.
a.
Cleaving off phosphate groups releases energy e.g. ATP
b
involved as coenzymes in energy-transferring reactions e.g. NAD
Nucleotide coenzymes
Coenzymes are nonprotein substances that are required for enzyme action.
Coenzymes usually are adenosine nucleotides combined with vitamins of B complex.
Nicotinamide is B vitamin niacin.
NAD
NADP
FAD
Coenzyme - A
=
=
=
=
nicotinamide adenine dinucleotide (Karp: p. 110 Fig. 3.27)
nicotinamide adenine dinucleotide phosphate
flavin adenine dinucleotide
ATP plus B vitamine pantothenic acid
1e 5) Ribonucleic acid (RNA) (Fig. 2.53b)
Chain of ribonucleotides.
Joined by 3'-5' phosphodiester linkage or bond.
Sugar phosphate backbone:
Phosphate atom is esterified to 2 oxygen atoms on adjoining sugars
Sugar is always ribose.
Base can be:
1. Adenine
2. Guanine
3. Cytosine
4. Uracil
but NOT thymine
Usually single stranded:
one end is the 5' end.
the other end is the 3' end
BIOL 130
MODULE 1
1e 6) Deoxyribonucleic acid (DNA) (Karp: pp. 386-388 Figs. 10.9, 10.10)
Chain of deoxyribonucleotides.
Joined by 3'-5' phosphodiester linkage or bond.
Sugar phosphate backbone:
Phosphate atom is esterified to 2 oxygen atoms on adjoining sugars
Sugar is always deoxyribose.
Base can be:
1. adenine
2. guanine
3. cytosine
4. thymine
but NOT uracil
DNA is double stranded (Karp: p. 388 Fig. 10.10)
Double-stranded helix with the two strands lying antiparallel
Two strands are held together by hydrogen bonds between bases.
Hydrogen bonds are between complementary pairs of purines and pyrimidines.
Rules of base pairing:
Adenine pairs with Thymine (AT)
Guanine pairs with Cytosine (GC)
17
BIOL 130
MODULE 1
1f: Proteins: Unit #6
(Karp: pp. 49-63)
1f 1) Protein Functions
Proteins in brief
Consist of one or more polypeptide chains.
A polypeptide is a polymer of amino acids linked together by peptide bonds.
Our cells may have as many as 100,000 different proteins.
Proteins have a diverse array of functions.
Some general functions of proteins
1.
2.
3.
4.
5.
6.
7.
8.
Enzymes protein catalysts
Structural elements e.g. tubulin
Contractile elements e.g. myosin
Control activity of genes e.g. transcription factors
Transport material across membranes e.g. glucose transporter
Carriers e.g. hemoglobin
Hormones e.g. insulin
Antibodies
1f 2) Amino Acids (Figs. 2.24, 2.26)
Building blocks of proteins.
Organic acids that contain an amino group
general structure
H H
H N C C OH
R
amino group
carboxyl group
carbon = first carbon after the carboxyl group
18
BIOL 130
MODULE 1
19
R groups Fig 2.26
R
= side chains of variable structure.
= any of 20 different groups
Differences in R groups accounted for different properties of amino acids and
proteins.
R groups can be broadly classified as:
1. Polar charged
2. Polar uncharged
3. Nonpolar
4. R groups with unique properties.
1f 3) Peptide bond or Amide linkage (Karp: p. 50 Fig. 2.24)
links -amino group of one amino acid with -carboxyl group of adjoining amino acid.
the linkage in dipeptides and in polypeptides.
R groups are not involved.
1f 4) Primary Structure of protein
The sequence of amino acids in a polypeptide.
Most polypeptides contain over 100 amino acids.
In a polypeptide chain, amino acids are residues.
N-terminus is the end of a polypeptide with a free amino group.
C-terminus is the end of a polypeptide with a free carboxyl group.
1f 5) Protein confirmation
Three dimensional structure of a protein.
Secondary, tertiary, and quaternary structure describes confirmation.
Primary structure determines secondary, tertiary, and quaternary structures of proteins.
1f 6) Secondary structure of protein
Results from hydrogen bonding between the oxygen of one peptide group and the
nitrogen of another peptide group.
Secondary, tertiary, and quaternary structure describes confirmation.
R groups are not involved.
Fixed configuration of the polypeptide backbone.
Secondary structure is limited to a small number of conformations.
Two common secondary structures.
BIOL 130
MODULE 1
20
helix (Karp: p. 55 Fig. 2.30)
Cylindrical, twisting spiral
Each amino acid is hydrogen bonded to its fourth neighbor on both sides.
pleated sheets or -pleated sheets (Karp: p. 55 Fig. 2.31)
Polypeptide chains of pleated sheets are stretched out and lie side by side, either
parallel or anti-parallel to one another.
Bonded groups may be portions of same chain folded back on itself or bonded
groups may be on separate chains.
Unorganized portions of protein
60% of the polypeptide chain in an average protein exists as helices and sheets.
Remainder is in random coils and turns.
Do not get continuous helix or pleated sheet for two reasons:
1. Juxtaposition of two bulky or similarly charged side chains.
2. Proline 'helix breaker'
Depiction of secondary structure (Karp: p. 56 Fig. 2.32)
helices are represented by helical ribbons.
strands as flattened arrows.
Unorganized connecting segments are thin strands in loops and turns.
1f 7) Tertiary Structure
The way that regions of secondary structure are oriented with respect to each other.
Tertiary structure predominates in globular proteins.
Monomeric proteins consist of a single polypeptide chain folded into its tertiary
structure.
Tertiary structure results from side chain interactions (Karp: p. 58 Fig. 2.35).
These are:
1. Hydrogen bonds
2. Hydrophobic bonds
3. Ionic bonds
4. Disulfide bond (Karp: p. 53)
covalent bond between two cysteines
BIOL 130
MODULE 1
21
1f 8) Motifs vs. Domains
Motifs (Karp: p. 510-511 Figs 12.37, 12.38)
A substructure found among many different proteins.
Recurring combinations of secondary structure.
Characteristic combinations of helices, sheets and loops found in a variety of proteins.
Usually associated with a particular function.
e.g. Zinc finger (see section 4c4 and Karp p 521)
Bundle of three secondary structures: an helix and a pair of anti-parallel
strands.
This motif generally is present in proteins that bind DNA.
Domains (Karp: p. 58 Fig. 2.36)
Found often in large proteins.
A region within a protein that folds and functions in a semi-independent manner.
Domains are modules of tertiary structure.
Different domains of a polypeptide often represent parts that function in a semi
independent manner. e.g. bind different factors
Some polypeptides containing more than 1 domain are thought to have arisen
during evolution by fusion of genes.
1f 9) Quaternary Structure (Karp: p. 60 Fig. 2.38)
Most proteins are made up of more than 1 polypeptide chain.
Each chain is a subunit.
A protein with 2 identical subunits = homodimer
A protein with 2 non-identical subunits heterodimer
Often the subunits can be independently folded.
Quaternary structure is the spatial arrangement of these subunits.
The bonds involved are the same as in tertiary structure.
1f 10) Covalent Modifications of Proteins
Modifications made by additions to the amino acid R groups.
An example is placement of a phosphate on serine, threonine and tyrosine residues.
(Karp p. 607; Fig. 15.3)
BIOL 130
MODULE 1
22
1f 11) Other structural Features of Proteins
Fibrous vs. globular proteins
On basis of overall confirmation.
Fibrous:
Has highly elongated shape.
Found outside living cells.
Globular:
A compact shape.
Most proteins within a cell are globular.
1f 12) Multiprotein complexes (Karp: p. 61 Fig. 2.41)
Physical association of different proteins, each with a specific function, to coordinate
a larger function.
e.g. pyruvate dehydrogenase complex
BIOL 130
MODULE 2
Module 2 outline and notes: Enzymes and energy metabolism
2a Bioenergetics
2a1) Introduction to Bioenergetics
2a2) The Laws of Thermodynamics
2a3) Free Energy
2a4) ATP and Coupled Reactions
2a5) Oxidation-reductions and making of ATP
2a6) Three General Processes of ATP Formation
2a7) Utilization of Free Energy of ATP
2b Enzymes
2b1) Introduction to Enzymes
2b2) Active site and Molecular Specificity
2b3) Denaturation and enzyme activity
2b4) Enzyme inhibitors
2b5) Naming enzymes
2b6) Metabolic regulation
2b7) Feedback principle
2c Metabolism
2c1) Introduction to Metabolism
2c2) Glycolysis
2c3) Reducing Power
2c4) Equilibrium vs. Steady State Metabolism and Regulation
2c5) Separating Catabolic and Anabolic Pathways
2c6) Separating and regulating glycolysis and gluconeogenesis
2c7) Regulating ATP levels
23
BIOL 130
MODULE 2
24
2a: Bioenergetics: Unit #7
(Karp: pp. 84-92; 107; 182; 185-188)
2a 1) Bioenergetics
Study of various types of energy transformations that occur in living organisms.
Energy:
Capacity to do work.
Work is to change or move something
Thermodynamics:
Study of changes in energy that accompany events in the universe.
Predict direction that events will take.
Predict whether or not an input of energy is required to cause the event to happen.
2a 2) The Laws of Thermodynamics
First Law of Thermodynamics
Energy can neither be created nor destroyed.
However, energy can be converted from one form to another.
e.g. conversion of sunlight into chemical energy
Second Law of Thermodynamics
Events in the universe have direction.
Events proceed from a state of higher energy to a state of lower energy.
Such events are spontaneous. They are thermodynamically favorable and can occur
without input of external energy.
The amount of useable energy is reduced.
2a 3) Free energy ( G)
1.
2.
3.
G is a change during a process in the energy available to do work.
A negative free energy ( G) means that the energy transformation may proceed
spontaneously.
a.
This is thermodynamically favorable and is an exergonic process.
b. Energy is available for use in another process.
A positive free energy (+ G) means that the energy transformation may not
proceed.
a.
This is thermodynamically unfavorable and is an endergonic process.
b. Energy must be added for process to proceed.
BIOL 130
MODULE 2
25
Standard-free energy difference ( G )
Describes difference in free energy when one mole of each reactant is converted to
one mole of each product at 25C and 1 atm of pressure.
Free-energy changes in metabolic reactions
Cells can carry out many reactions with positive G .
1.
2.
Ratio of the reactant to product is maintained above that defined by equilibrium
constant by other cellular reactions.
Couple endergonic and exergonic reactions.
Coupling occurs when an endergonic reaction and an exergonic reaction are
catalyzed by same enzyme.
2a 4) ATP and coupled reactions (Karp: pp. 90-91)
Link between exergonic and endergonic reactions is ATP.
Hydrolysis of ATP drives endergonic reactions.
ATP + H2O ADP + phosphate G = -7.3 kcal/mole
glutamic acid + NH3 glutamine + H2O G = 3.4 kcal/mole
glutamic acid + ATP + NH3 glutamine + ADP + HPO G = -3.9 Kcal/mole
ATP acts as a common intermediate in biological-energy transfer.
2a 5) Oxidation reduction (or redox) reactions and making of ATP
Loss of Electrons from a substance is Oxidation (LEO).
Gain of Electrons by a substance is Reduction (GER).
Oxidations and reductions always occur together.
Substance that is oxidized (donates electrons) is reducing agent.
Substance that is reduced (gains electrons) is the oxidizing agent.
Oxidation state of carbon atom (Karp: p. 107 Fig. 3.23)
1.
2.
3.
Is not obvious but here are 2 helpful rules.
a.
Oxidized = O2 added or hydrogen removed.
b. Reduced = O2 removed or hydrogen added.
Degree of reduction of a compound is a measure of its ability to perform chemical
work within the cell.
Carbohydrates are rich in chemical energy because they contain strings of
|
H-C-OH
|
e.g. glucose
BIOL 130
MODULE 2
26
Oxidation-Reduction potential (Karp: pp. 106; 182, 186, Fig 5.14)
1. The tendency of a substance to accept or to donate electrons in a chemical reaction
with another substance is defined by its oxidation-reduction potential or redox
potential.
2. This is expressed in volts.
_________________________________________________
-0.6
-0.4
-0.2
0
+0.2
+0.4
+0.6
3. When compound has a negative value, it readily donates electrons.
4. When compound has a positive value, it readily accepts electrons.
5. Electrons can be transferred spontaneously from a compound to another compound
that has a more positive redox potential.
6. The greater the gap in redox potential between the donor and the acceptor, the
greater the standard free-energy change.
7. This energy can be used for the phosphorylation of ADP to ATP.
Electron-transport chain
1.
2.
3.
4.
5.
6.
Electron carriers are capable of existing in either an oxidized or reduced state.
Electrons are passed from 1 carrier to next until final acceptor becomes reduced.
Each electron carrier along chain has a more positive redox potential than previous one.
With each successive transfer, electrons loose additional free energy.
Free energy released by electron transfer is utilized to generate a proton gradient.
Proton gradient is utilized to drive formation of ATP.
2a 6) Three General Processes of ATP Formation
Oxidative phosphorylation
Formation of ATP in electron transport chain of mitochondria.
Photophosphorylation
Formation of ATP in electron transport chain of chloroplast.
Substrate-level phosphorylation (Fig. 3.28)
ATP formation is coupled to the hydrolysis of phosphate compounds with a higher Go'.
e.g. glycolysis
2a 7) Utilization of free energy of ATP (Karp: p. 91-92 Fig. 3.6; Fig. 3.7)
1.
2.
3.
4.
Separate charges across a membrane. e.g. function of nerves and muscle
Concentrate a particular solute within the cell e.g. active transport
Drive an otherwise unfavorable reaction e.g. protein synthesis
Slide filaments across one another. e.g. cell movement
BIOL 130
MODULE 2
27
2b: Enzymes Unit #8
(Karp: p. 92-103; 112-116)
2b 1) Introduction to Enzymes
Catalysts
1.
2.
3.
4.
Cell is a collection of chemical reactions that together allow a cell to persist.
Cell controls chemical reactions by using catalysts.
Catalysts speed up rate of a reaction but are unchanged themselves
Catalysts made by cells are:
a.
Enzymes (most cell catalysts) protein catalysts
Enzymes are actually proteins plus nonprotein components, cofactors.
Cofactors may be inorganic (metals) or organic (coenzymes)
b. ribozyme (rare) RNA catalysts
Some RNAs catalyze RNA cleavage.
Reactants
1. Reactants bound by an enzyme are called substrates.
2. Substrates are converted to products.
3. Rate of a reaction is proportional to concentration of reactants.
4. All chemical reactions proceed toward a state of equilibrium.
5. At equilibrium, rates of forward and backward reactions are equal.
6. Ratio of products to reactants at equilibrium is an inherent property of reacting
chemicals.
7. The direction that the reaction is proceeding at any moment is dependent on relative
concentrations of all molecules at that moment.
Properties of enzymes
1.
2.
3.
4.
5.
Enzymes are effective in small amounts.
Enzymes are unchanged by reactions that they catalyze.
Enzymes have specificity.
They catalyze only one or a small number of closely related reactions.
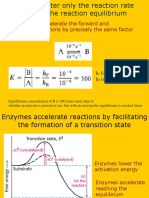
Enzymes do not change the direction of reactions
Only rate at which reactions reach equilibrium.
Enzymes only accelerate rate at which a favorable chemical reaction proceeds.
(Table 3.3)
Enzymes have no effect on thermodynamics of reaction.
BIOL 130
MODULE 2
Overcoming Activation Energy Barrier
1.
2.
3.
4.
Even thermodynamically favorable reactions do not proceed on their own at
relatively rapid rates in absence of enzymes ( p. 93 Table 3.3).
e.g. glucose is kinetically stable even if thermodynamically unstable.
Chemical reactions require that certain covalent bonds be broken within reactants.
Reactants must contain sufficient kinetic energy that they overcome a barrier,
activation energy. (Fig. 3.8)
enzymes catalyze reactions by decreasing magnitude of activation energy barrier.
2b 2) Active site and Molecular Specificity (Figs. 3.10, 3.11, 3.12, 3.14)
Enzymes:
Accelerate bond-breaking and bond-forming processes.
Do this by forming a complex with reactants (enzyme-substrate complex).
Bind substrates by noncovalent interactions at active site.
Active or catalytic site of enzymes:
Part of enzyme directly involved in binding substrate.
Typically buried in cleft or crevice of the enzyme.
Accounts for specificity of enzymes.
Each enzyme exhibits a very great specificity for a particular reaction.
2b 3) Denaturation and enzyme activity (Karp: p. 62 Fig. 2.43)
Denaturation is unfolding or disorganization of a protein.
Destroying secondary and higher structure is called denaturation.
Specificity and catalytic properties of enzymes are a result of their 3 dimensional
structure.
Therefore, denaturation destroys enzyme activity.
2b 4) Enzyme Inhibitors (Karp: pp. 102-103)
Molecules that bind to an enzyme and decrease its activity.
Can be of two types: irreversible or reversible.
Irreversible inhibitors
Bind very tightly to an enzyme
Often form a covalent bond to one of enzyme's amino acid residues.
Often man made reagents. e.g. organophosphate pesticides
Made in nature as part of chemical warfare. e.g. penicillin
28
BIOL 130
MODULE 2
Reversible inhibitors
Bind loosely to an enzyme and can be readily displaced.
Can be of two types.
Competitive inhibitors (Karp: p. 102 Fig. 3.20)
Compete with substrate for active site.
Structurally resemble substrate
But cannot be transformed into product.
Can be overcome if substrate/inhibitor ratio is great enough.
Noncompetitive inhibitors
Does not bind at same site as substrate.
Inhibitor acts at a site other than active site.
Level of inhibition depends only on concentration of inhibitor.
Concentration of substrate cannot overcome it (Karp: p. 104 Fig. 3.21a)
2b 5) Naming enzymes
Informal or trivial names
Named by tacking -ase onto end of substrate
A few hints.
Kinase usually involves ATP
e.g. hexokinase and pyruvate kinase
dehydrogenase usually involves coenzyme NADH
e.g. glyceraldehyde phosphate dehydrogenase
some exceptions to -ase ending
e.g. trypsin and pepsin
2b 6) Metabolic Regulation (Karp: pp. 112-116)
Enzymes are regulatory elements in metabolism.
Metabolism is sum total of all chemical changes in cells.
Metabolism can be controlled by:
1. Controlling concentrations of enzymes (long term control)
discussed later under gene regulation
2. Changing an enzyme's activity
Two important ways:
a.
Covalent modification
b. Allosteric modulation
29
BIOL 130
MODULE 2
30
Covalent modification (Karp: p. 112-113; p. 607 Fig 15.3; p. 619 Fig. 15.12)
1.
2.
3.
Many enzymes exist in active and inactive forms.
e.g. glycogen phosphorylase (gp)
glycogen phosphorylase a active
glycogen phosphorylase b inactive
Active and inactive forms are interconverted by covalent modifications.
Addition of 1 phosphate on a specific serine residue makes gp active.
Removal of this phosphate makes gp inactive.
These are catalyzed by other enzymes.
Protein kinases puts phosphate groups onto proteins.
Protein phosphatases removes phosphates from proteins.
Allosteric modulation (Karp: p. 113)
1.
2.
3.
4.
5.
Activity is modulated through noncovalent binding of a specific metabolite at a site
on the protein other than active site.
There can be negative or positive modulators.
They change the conformation of the enzyme.
Positive modulators stabilize a conformation that has a high affinity for substrate.
Negative modulators stabilize a conformation that has a low affinity for substrate.
2b 7) Feedback principle (Karp: p. 113 Fig. 3.30)
1.
2.
3.
4.
Allosteric enzymes are often first enzyme in a metabolic pathway.
They are subject to feedback inhibition.
intracellular level of a substance regulates the rate of its own synthesis.
Controls activity of one or more enzymes within pathway that produces it.
E1
E2
E3
BIOL 130
MODULE 2
31
2c: Metabolism: Unit #9
(Karp: pp. 105-116)
2c 1) Introduction to Metabolism
Sum total of all chemical changes that occur in cells.
Made up of interconnected metabolic pathways.
Pathways are sequence of chemical reactions.
Each reaction is catalyzed by a specific enzyme.
Compounds formed in each step along the pathway are metabolites.
Pathway leads to an endproduct.
Endproduct has a particular role in the cell.
Two broad types of metabolic pathways.
Catabolic pathways
Breaking of chemical bonds in large, complex molecules to form small simple molecules.
These molecules can be used to synthesize other molecules.
Catabolic reactions are exergonic.
Provide chemical energy for cell
Anabolic pathways
Synthesis of large molecules by chemically bonding together small molecules.
Anabolic reactions are endergonic.
Require energy.
The two are interconnected.
Catabolic pathways provide energy and small molecules for anabolic pathways.
2c 2)Glycolysis (Karp: p. 108 Fig. 3.24)
1.
2.
3.
4.
5.
A universal catabolic pathway.
Breakdown of glucose.
Ten-step reaction sequence (glucose to pyruvate).
Occurs in presence or absence of O2.
Occurs in cytosol.
Glycolytic products for 1 molecule of glucose
1.
2.
Uses 2 ATP and produces 4 ATP
ATP formed by substrate-level phosphorylation
Yields 2 pyruvates
pyruvate stands at junction of anaerobic vs. aerobic
BIOL 130
3.
MODULE 2
Yields 2 NADH
a.
Fermentation
b. Electron transport chain in mitochondria
c.
Make NADPH
2c 3) Reducing Power
1.
2.
3.
4.
5.
6.
A cell's reservoir of NADPH.
Formation of complex biological molecules requires reduction of precursors.
Accomplished by transfer of high-energy electrons from NADPH.
NADPH is coenzyme for enzymes having reductive role in anabolic pathways.
e.g. C3 cycle in photosynthesis
e.g. fat
NADPH is interconvertible with NAD+.
NAD+ is coenzyme for dehydrogenases in catabolic pathways.
e.g. glyceraldehyde dehydrogenase in glycolysis
2c 4) Equilibrium versus steady state metabolism (Karp: p. 92 Fig. 3.7)
Many reactions in a metabolic pathway may be near equilibrium.
However, several reactions in a pathway are poised far from equilibrium.
These are essentially irreversible and keep pathway going in a single direction.
Cellular metabolism can maintain itself at irreversible nonequilibrium conditions
because the cell is an open system.
An open system because materials are continually flowing in and out of the cell.
Cellular metabolism is said to exist in a steady state.
In a steady state, the concentrations of reactants and products remain essentially
constant, even though individual reactions are not necessarily at equilibrium.
Driving force for glycolysis
When concentration of metabolites in the cell are measured,
Three reactions are far from equilibrium (Figs. 3.24, 3.25, 3.31):
1. Hexokinase
2. Phosphofructokinase
3. Pyruvate kinase
These 3 are essentially irreversible and keep pathway going in a single direction.
All subject to feedback inhibition by ATP.
32
BIOL 130
MODULE 2
33
2c 5) Separating Catabolic and Anabolic Pathways
1.
2.
3.
e.g. glycolysis and gluconeogensis
Glycolysis (catabolic) breakdown of glucose
Gluconeogenesis (anabolic) formation of glucose
Thermodynamic problem cannot proceed simply by reversal of reactions.
Glycolytic pathway contains 3 thermodynamically irreversible reactions. (Fig. 3.31)
Regulatory problem two pathways could not be controlled independently of one
another.
2c 6) Separating and regulating glycolysis and gluconeogenesis (Fig. 3.31)
1.
2.
Solved by using different enzymes to catalyze 3 key reactions in 2 opposing pathways.
e.g. Step between fructose-6-phosphate and fructose 1,6-bisphosphate
Other reactions are identical, although they run in opposite directions.
Phosphofructokinase in glycolysis
fructose 6-phosphate + ATP ADP + fructose 1, 6-bisphosphate
This is an allosteric enzyme.
positive modulators: ADP, AMP, fructose 2, 6-bisphosphate
negative modulator: ATP
When cell has ample ATP, the enzyme is inhibited.
If ATP is being used up, the enzyme is stimulated.
Fructose 1,6-bisphosphatase in gluconeogenesis
fructose 1,6-bisphosphate + H2O fructose 6-phosphate + pi
Allosterically inhibited by high levels of AMP.
Competitively inhibited by fructose 2, 6-bisphosphate.
2c 7) Regulating ATP levels
Generally ATP levels do not fluctuate
ATP must be maintained high relative to that of ADP and AMP.
In this way, - G of ATP hydrolysis remains large enough to drive endergonic reactions.
BIOL 130
MODULE 2
34
Here are some sample midterm questions.
1. Which one of the following specific compounds is correctly matched with the general
class of compound to which it belongs?
A. insulin / carbohydrates
B. testosterone / protein
C. cellulose / nucleic acid
D. insulin / lipid
E. estrogen / lipid
2. Which one of the following molecules would be considered macromolecules?
A. polypeptides
B. organic acids
C. purines
D. hexoses
E. oligosaccharides
3. Which one of the following statements about enzymes is not true?
A. Enzymes are proteins.
B. Enzymes regulate metabolism.
C. Enzymes change the direction of a chemical reaction.
D. Enzymes can be covalently modified.
E. If all (A to D) are true, answer E.
4. What is the repeating disaccharide in cellulose?
A. maltose
B. cellobiose
C. lactose
D. sucrose
E. none of the above
5. Which one of the following completions is correct? Eukaryotic cells . . .
A. include only cells that have cell walls.
B. have their DNA organized into a nucleus.
C. have no organelles.
D. have their DNA organized into a nucleoid region.
E. are found only in multicellular organisms.
BIOL 130
MODULE 2
35
6. Which one of the following statements about cellular metabolism is true?
A. Cellular metabolism exists in equilibrium.
B. All steps in a metabolic pathway are irreversible.
C. One or more reactions in a metabolic pathway are far from equilibrium.
D. Irreversible inhibitors bind to an enzyme through weak bonds.
E. If none of the above statements are true, answer e.
7. Which one of the following completions is NOT true?
Human cells ..............
A. always are eukaryotic cells.
B. always contain a cell membrane.
C. always lack cell walls.
D. are one component of human tissues.
E. never have a nucleus.
8. Which one of the following completions is correct? Microtubules are..............
A. 20 to 25 nm in diameter and found only in prokaryotes.
B. 10 nm in diameter and found in plant, animal and bacterial cells.
C. 5 nm in diameter and found only in eukaryotes.
D. 20 to 25 nm in diameter and found in plant and animal cells.
E. 5 nm in diameter and found in plant, animal and bacterial cells.
For the next 2 questions refer to the figure below (not produced here but look at Figure
2.30 b & c in Karp).
9 Which one of the following statements about the above figure is correct?
A. The figures represent a double helix, which is a characteristic structure of DNA.
B. The figures represent an alpha helix, which is an organization frequently found in
RNA.
C. The figures represent an alpha helix, which is found in nutritional polysaccharides.
D. The figures represent a pleated sheet, which is an organization frequently found in
proteins
E. The figures represent an alpha helix, which is an organization frequently found in
proteins
10. In the above figure, what type of bond maintains the structure?
A. A hydrogen bond between R groups.
B. A hydrogen bond, which is a weak bond.
C. A hydrophobic bond, which is a weak bond.
D. A hydrophobic bond between R groups.
E. An ionic bond between peptide bonds.
BIOL 130
MODULE 2
ANSWERS TO SAMPLE MIDTERM QUESTIONS on previous page.
1. E
2. A
3. C
4. B
5. B
6. C
7. E
8. D
9. E
10. B
36
BIOL 130
MODULE 3
Module 3 outline and notes: Membranes and energy metabolism
3a Membranes
3a1) Introduction to Membranes
3a2) Overview of Plasma Membrane Structure
3a3) Membrane Lipids
3a4) Plasma Membrane Carbohydrates
3a5) Membrane Proteins
3a6) Motility of Membrane Proteins and Cell Polarity
3a7) Specialized membrane structures: intercellular junctions
3b Membrane transport
3b1) Movement of Substances Across Plasma Membranes
3b2) Diffusion
3b3) Solute Transport Mechanisms
3b4) Diffusion through Lipid Bilayer
3b5) Diffusion of Ions Through Membranes
3b6) Facilitated Diffusion
3b7) Active Transport
3b8) Na+-K+ ATPase
3b9) Coupling Active Transport to Existing Ion Gradients
3b10) Membrane Potentials
3c Cellular uptake of particles and macromolecules
3c1) Introduction to Cellular Uptake of Particles and Macromolecules
3c2) Endocytosis
3c3) Pathways in receptor-mediated endocytosis
3c4) LDLs and cholesterol metabolism
3c5) LDL and HDL in heart disease
3c6) Phagocytosis
3c7) Some bacteria circumvent being killed by macrophages
3d ATP formation and the mitochondria
3d1) Fermentation: Anaerobic Oxidation of Pyruvate
3d2) Aerobic Oxidation of Pyruvate
3d3) TCA or Krebs Cycle
3d4) Mitochondria Structure
3d5) Electron Transport Chain
3d6) ATP formation
3d7) Overall Products from oxidizing glucose
3e Photosynthesis and the chloroplast
3e1) Heterotrophs vs. Autotrophs
3e2) Chloroplasts
3e3) Absorption of Light
3e4) Reaction-Center Chlorophyll
3e5) Photophosphorylation
3e6) CO2 Fixation and Formation of Carbohydrates
37
BIOL 130
MODULE 3
38
3a: Membranes: Unit #10
(Karp: p. 117-141; 250-258)
3a1) Introduction to Membranes
Continuous, unbroken sheets, enclosing compartments.
Dynamic structures capable of fusing without loosing continuity.
Plasma membrane: encloses contents of entire cells.
Nuclear envelope
Mitochondrial membranes
Chloroplast membranes
Lysosomal membrane
Endoplasmic reticulum
Summary of Membrane Functions
1. Compartmentalization
2. Provide a selectively permeable membrane
3. Transporting solutes
4. Responding to external signals
signal transduction
5. Intercellular interaction
6. Locus for biochemical activities
7. Energy transduction
3a2) Overview of plasma membrane structure
1.
2.
Biochemistry
a.
Consists of polar lipids arranged in a bilayer.
b. Proteins
c.
Carbohydrates
Appearance of plasma membrane in electron microscope
a.
Thin sectioning technique
~7.5 nm wide with dark, light, dark appearance (Karp: p. 118 Fig. 4.1)
b. Freeze fracture technique
Smooth areas interrupted by bumps and depressions. (Karp: p. 129 Fig. 4.15)
Fluid mosaic model (Karp: p. 121 Fig. 4.4)
1.
2.
3.
4.
5.
6.
Lipid bilayer is core of membrane
Lipid molecules are present in a fluid state capable of rotating and moving laterally
within membrane.
Proteins occur as "mosaic" of discontinuous particles.
Some proteins penetrate deeply into, and even through, lipid bilayer.
Membranes are dynamic structures in which components are mobile.
Components come together to engage in various transient interactions.
BIOL 130
MODULE 3
39
3a3) Membrane Lipids
All are amphipathic.
Except for glycolipids and cholesterol, all contain phosphate group.
These are phospholipids.
Phospholipids have a glycerol backbone.
These are phosphoglycerides.
Phosphoglycerides (Karp: p. 48 Fig. 2.22; p. 123 Fig. 4.6)
Diglycerides often one saturated and one unsaturated fatty acid.
3rd OH group has phosphate plus either:
1.
2.
3.
4.
Choline phosphatidyl choline.
Ethanolamine phosphatidylethanolamine.
Serine phosphatidylserine.
Inositol phosphatidylinositol.
Sphingolipids (Karp: p. 123 Fig. 4.6b)
Sphingosine-based lipids.
Sphingosine is an amino alcohol with a long hydrocarbon chain.
Two additions:
1. Always a fatty acid to amino group of sphingosine. This is a ceramide.
2. Additional groups esterified to terminal OH.
a.
If phosphorylcholine, molecule is sphingomyelin.
b. If carbohydrate, molecule is glycolipid.
glycolipids
1.
2.
If carbohydrate is a monosaccharide, glycolipid is a cerebroside.
If carbohydrate is an oligosaccharide, glycolipid is a ganglioside.
Cholesterol (Karp: p. 124 Fig. 4.7)
May constitute up to 50% of lipid molecules in plasma membrane of certain animal
cells.
Is absent from plasma membranes of all bacterial cells.
Cholesterol increases fluidity of bilayer.
BIOL 130
MODULE 3
40
3a 4) Plasma membrane carbohydrates
All of carbohydrate faces outward into extracellular space.
Covalently linked to either protein or lipid.
glycoproteins (Karp: p. 126 Fig. 4.11; p. 127 Fig. 4.12)
Carbohydrate is present in short, branched oligosaccharides (<15 sugars per chain).
May be attached to R group of:
1. Asparagine (N-linked)
2. Serine and threonine (O-linked)
Play roles in cell interactions.
glycolipids (Karp: p. 127 Fig. 4.12)
May play role in certain infectious diseases.
Carbohydrates of glycolipids determine blood type
(A, B, AB, or O)
3a 5) Membrane proteins
thousands of different ones.
Asymetrically situated.
Properties of outer portion of membrane are very different from properties of inner
portion.
3 distinct classes based on their relationships to lipid bilayer. (Fig. 4.13)
BIOL 130
MODULE 3
41
1. Integral proteins
Penetrate into lipid bilayer.
Usually pass right through.
Have segments protruding into extracellular space.
Have segments protruding into cytoplasm.
Transmembrane segments
Pass through lipid bilayer.
Usually consist of nonpolar amino acids organized in an -helical conformation.
Some integral membrane proteins can form aqueous channel through lipid bilayer.
2. Peripheral membrane proteins
Located entirely outside of lipid bilayer.
Either on extracellular or cytoplasmic surface.
Associated with membrane surface by noncovalent bonds.
Weak electrostatic bonds to hydrophilic head groups of phospholipids or
hydrophilic portion of integral proteins.
Peripheral proteins on cytoplasmic surface function in transmembrane signal transduction.
3. Lipid-anchored membrane proteins
Covalently linked to lipid molecules within bilayer.
Two types of lipid anchors:
a.
Glycophosphatidyl inositol in outer leaflet
proteins linked to this by short oligosaccharide chain
b. Long hydrocarbon chains in inner leaflet
A G protein called Ras is linked this way.
G proteins are involved in transmembrane signal transduction.
3a6) Motility of Membrane Proteins and Cell Polarity
Control over membrane motility
1.
2.
3.
Integral membrane proteins are not totally free to drift in lipid sea.
Restraints as a result of:
a.
Interactions occurring within membrane.
b. Links to materials on inner or outer surface of membranes. (Karp: p. 142 Fig. 4.27)
Leads to membrane domains
Regions in which a protein may travel (Karp: p. 142 Fig. 4.27)
Membrane Domains and Cell polarity (Karp 140)
e.g. Epithelial cells lining inner surface of intestine. ( Fig. 4.30)
BIOL 130
MODULE 3
Highly polarized cells whose different surfaces carry out different functions.
1. Apical plasma membrane
Faces lumen of intestine
Selectively absorbs substances from lumen.
2. Lateral plasma membrane surface
Interacts with neighboring epithelial cells.
3. Basal plasma membrane
Adheres to underlying extracellular substrate (basement membrane)
Functions in exchange of substances with bloodstream.
3a7) Specialized membrane structures: intercellular junctions
Nonjuctional
Space between neighboring cells ~20-30 nm.
No specialized structures are seen in space or in cells.
Specialized cell-cell junctions
Specialized regions of close-range contact between adjacent cells.
1. Adherens junctions
2. Desmosomes
3. Tight junctions
4. Gap junctions
Often occur together as intercellular junctional complex.
Adherens Junctions and Desmosomes: anchoring cells to other cells
Adherens junctions (zonulae adherens) (Karp: p. 252; Fig. 7.25; p. 252 Fig. 7.26)
Sites where cadherins are concentrated.
Space between cells is 20-35 nm.
In cytoplasm, structures called plaques seen.
Cadherins connected to actin filaments by catenin.
Very common in epithelia, such as gut epithelium.
Occur as 'belt' that encircles each of cells near its apical surface
Binds that cell to its neighbors.
Desmosomes (Karp: p. 253 Fig. 7.27 & 7.28)
Sites where cadherins (desmogleins and desmocollins) are concentrated.
Space (desmoglea) between cells is 20-35 nm.
Desmolgea contains collagenous glue.
Opposite desmoglea are dense cytoplasmic plaques.
Intermediate filaments run out of plaques across cell.
Desmosome abundant in epithelia.
42
BIOL 130
MODULE 3
Occur basal to adherens junctions.
'spot welding'.
Tight Junctions (TJ): sealing extracellular space (Karp: p. 254 Fig. 7.30)
No space between two neighboring epithelial cells.
Two membranes together are ~12 nm thick rather than ~14 nm.
Appear to involve fusion of outer leaflets of adjacent membranes
5 layered structure as seen in TEM.
Claudins are the proteins of tight junctions.
TJ occur at very apical end of junctional complex
Serve to seal space between cells.
In intestine, tight junctions form a ring or zone completely around each cell.
This forces material to go through cells rather than between cells.
Gap junctions: mediate cell-to-cell communication (Karp: p. 257; Fig. 7.32; 7.33)
Space between adjacent cells is 2-4 nm.
7 layered structure as seen in TEM.
Extremely small channels (connexons) span the gap.
Connexons allow for cytoplasmic continuity between adjacent cells.
Only low molecular weight material can pass between cells.
Gap junctions act as molecular sieves.
Integrate activities of individual cells of a tissue.
43
BIOL 130
MODULE 3
44
3b: Membrane Transport: Unit #11
(Karp: p. 143-155; 158-161)
3b1) Movement of substances across plasma membranes
1.
2.
Plasma membrane is selectively permeable.
a
i.e. some solutes are more permeable than others.
b. This allows cells to concentrate substances
c.
Influx movement of substance into cells.
d. Efflux movement of substances out of cells.
e.
Net flux one exceeds the other.
Plasma membrane is semipermeable.
a.
i.e. freely permeable to H2O (solvent), while allowing much slower passage to
small ions and polar solutes.
3b2) Diffusion
Continual movement of molecules among each other in liquids or gases
Greater the concentration difference the greater is the rate of diffusion.
Ability of an ion to diffuse between two compartments depends on 2 gradients:
1. Chemical gradient determined by concentration difference of substance.
2. Electric potential gradient determined by difference in charge.
3. Together these differences are combined to form an electrochemical gradient.
Diffusion of Water through Membranes
Osmosis Is movement of water through a semi-permeable membrane in response to a
concentration gradient of H2O.
Osmotic pressure is proportional to the number of particles in solution
Osmolarity is a calculated quantity of a solution.
Units are milliosmoles.
Tonicity Is an observed property that depends on permeability of the cell membrane
and is relative to another cell or solution (Figs. 4.35).
1.
If cell swells, the solution is hypotonic relative to cell.
a.
animal cells swell and eventually burst.
If these are red blood cells, this is hemolysis.
b. plant cells push against surrounding cell wall.
This is turgor pressure. (Karp: p. 146 Fig. 4.36)
2.
If cell shrinks, the solution is hypertonic relative to cell.
Plant cell pulls away from surrounding cell wall.
This is plasmolysis.
If cell neither shrinks or swells, the solution is isotonic.
3.
BIOL 130
MODULE 3
45
3b3) Solute transport mechanisms (Karp: p. 143 Fig. 4.33)
1.
2.
Passive diffusion
All driven by concentration gradient.
a.
Simple diffusion through lipid bilayer
b. Simple diffusion through an aqueous channel
c.
Facilitated diffusion solute binds specifically to a membrane protein carrier.
Active energy coupled transport
Uses protein carrier but driven by ATP hydrolysis.
3b4) Diffusion through Lipid Bilayer
1.
Greater the lipid solubility, faster the diffusion into the cell. (Karp: p. 145 Fig. 4.34)
Partition coefficient is a measure of lipid solubility or hydrophobicity or nonpolarity.
partition coefficient = concentration in oil
concentration in H2O
2.
The smaller the molecule, the greater is the rate of diffusion.
Very small, uncharged molecules penetrate very rapidly.
e.g. O2 & CO2 appear to slip between adjacent phospholipids
3.
Ions and lager polar molecules, such as sugars and amino acids, cannot diffuse
through lipid bilayer.
3b5) Diffusion of ions through membranes
Ion channels:
Permeable to specific ions.
Each formed by integral membrane proteins that surround an aqueous pore.
Highly selective
Only one type of ion passes through pore.
Bidirectional.
Net flux depends on electrochemical gradient.
Gated channels
1. Most ion channels exist in either an open or a closed conformation.
2. Such channels are termed gated.
3. Voltage-gated channels
Conformation state depends on difference in ionic charge on two sides of membrane.
e.g. K+ channel (Karp: p. 150-151 Fig. 4.41; Fig. 4.43)
4. Chemical-gated channels
Conformational state depends upon binding of a particular substance.
e.g. Acetylcholine (neurotransmitter) acts on outer surface of certain cation channels.
BIOL 130
MODULE 3
46
3b6) Facilitated diffusion
Selective binding of substance to a membrane-spanning protein, a facilitative
transporter, improves permeability of a substance without altering direction in
which substance would tend to move on its own.
Does not require ATP.
Movement facilitated equally well in both directions.
Direction depends on concentration gradient.
Facilitated diffusion mechanism (Karp: p. 152 Fig. 4.44)
1. Solute binds to facilitative transporter on one side of membrane.
2. Triggers a conformation change in protein.
3. Solute is exposed to other surface of membrane.
4. Solute can then diffuse down its concentration gradient.
Glucose transporter: An example of facilitated diffusion.
Certain similarities between facilitative transporters and enzymes
1.
2.
3.
Both specific for molecules they transport.
Both have their activities subject to regulation.
Both exhibit saturation-type kinetics. (Karp: p. 152 Fig. 4.45)
3b7) Active transport
Selective binding of substance to a membrane-spanning protein, an active transporter,
allows passage of substance through membrane against electrochemical gradient for
that substance.
Active transporter acts by undergoing conformational change upon binding substance.
Substance is moved against a concentration gradient.
Movement occurs in only 1 direction.
Requires ATP or other form of energy such as flow of other substances down a gradient.
An example is Na+-K+ ATPase or sodium-potassium pump
3b8) Na+-K+ ATPase
3 Na+ pumped out to 2 K+ pumped in for each ATP consumed.
Electrogenic pump.
Contribute directly to separation of charge across membrane.
Na+ gradient provides a means by which free energy can be stored in cell.
This potential energy in ionic gradient can be utilized in various ways.
BIOL 130
MODULE 3
Na+-K+ ATPase transport cycle (Karp: p. 153 Fig. 4.46)
1.
2.
3.
4.
5.
6.
7.
8
Na+ ions bind to protein on inside of membrane.
ATP is hydrolyzed.
Conformation of Na+-K+ ATPase is changed.
Na+ ions are expelled to external space.
K+ ions bind on outside of membrane.
Phosphate group on protein is removed.
Protein snaps back to its original conformation.
K+ ions move to inside of cell.
3b9) Coupling active transport to existing ion gradients
1.
2.
3.
4.
5.
6.
7.
8.
Na+that has been pumped out is driven back in by concentration gradient.
Membrane almost impermeable to Na+.
Therefore transport protein is required.
Transport protein does not work unless it binds another molecule (e.g. glucose).
Glucose is driven into cell against its concentration gradient.
This is cotransport and driven by secondary active transport.
Symport two solutes are moved in same direction.
(i.e. glucose and Na+)
Antiport two solutes moved in opposite direction.
(i.e. Na+ and H+)
Secondary transport in intestine (Karp: p. 140 Fig. 4.30; p. 158 Fig. 4.49)
Uptake of glucose by cotransport occurs at apical plasma membrane.
Glucose diffuses through cytoplasm of intestinal epithelial cell.
At basal plasma membrane glucose is transported out of cell and into bloodstream
by facilitated diffusion.
3b10) Membrane potentials
All cells have a voltage gradient across their membrane or a membrane potential.
By convention, inside is negative with respect to outside (Fig. 4.51 & 4.52).
Magnitude varies between -15 and -100 mV.
Membrane potential arises from:
1. Passive properties of the membrane (semi-permeable)
2. Active transport in Na+/K+ ATPase
47
BIOL 130
MODULE 3
48
3c: Cellular Uptake of Macromolecules and Particles: Unit #12
(Karp: pp. 301-309)
3c 1) Introduction to Cellular Uptake of macromolecules and particles
How cells take in materials too large to penetrate plasma membrane.
Uptake of extracellular materials into cytoplasmic vesicles without breaking
continuity of plasma membrane.
Two different mechanisms:
1. Endocytosis uptake of fluid, dissolved solutes and suspended macromolecules.
2. Phagocytosis uptake of particulate matter. > 0.5 m
3c 2) Endocytosis
Can be divided into 2 categories.
a.
Bulk-phase endocytosis
Brings about uptake of extracellular fluids without recognition by surface of
plasma membrane.
Any molecules that happen to be present in enclosed fluid gain entry into
cells.
Occurs in a continual manner in many types of cells.
b.
Receptor-mediated endocytosis (RME)
Brings about uptake of specific macromolecules (ligands) following their
binding to receptors on plasma membrane.
(ligand any molecule that can bind a receptor)
Material taken up by endocytosis is delivered to a network of tubules and
vesicles.
These are collectively known as endosomes.
Two main types of receptors in receptor-mediated endocytosis (RME)
(Fig. 8.42)
1. house keeping receptors
- often bind nutrients and deliver them to cytoplasm
e.g. LDL receptor
2. signal receptors
- binds ligands that are hormones or growth factors.
- endocytosis of these leads to destruction of receptor.
- a process called "receptor down regulation".
Endosomes
Network of cytoplasmic membrane vesicles and tubules.
Early endosomes are located near peripheral region of cell.
(act as first sorting station)
Late endosomes are in interior part of cell. (main sorting station)
Late endosomes receive material from:
a.
Early endosomes
b. Golgi apparatus
Late endosomes can be thought of as a prelysosomal compartment.
BIOL 130
MODULE 3
49
Coated pits (Karp: p. 302-304 Figs. 8.37 & 8.38)
Sites on membrane where receptors for receptor-mediated endocytosis are
concentrated.
Surface is indented.
Indentation is covered on its cytoplasmic side by a bristly electron-dense material.
Proteins of the coated pits (Figs. 8.39, 8.40 & 8.41)
1. clathrin
- main structural protein
- composed of 3 heavy and 3 light chains organized into a triskelion.
- forms scaffold for the pit or cage or basket.
2. adpatins
-link ligand to clathrin cage
-adaptins organize into adaptors.
Adaptors bind
1. Clathrin on one side and
2. Cytoplasmic tails of specific membrane receptors on other side.
3. dynamin
-large GTP-binding protein.
-releases clathrin-coated vesicle from the membrane.
3c3) Pathways in receptor-mediated endocytosis (Karp: p. 306 Fig. 8.42)
1. common steps
a. Clathrin coat is removed from coated vesicle to yield uncoated vesicle.
b. Uncoated vesicle fuses with early endosome.
2. house keeping receptors.
a. ligand and receptor are separated from each other in early endosome.
b. Receptors bud off into recycling vesicles to be taken back to membrane.
c. Endosomal carrier vesicle buds off with ligands and fuses with late endosome.
d. From late endosome, ligand moves to lysosome for ultimate delivery to
cytoplasm.
3. signaling receptors
a. ligand and receptor bud off into endosomal carrier vesicle.
b. Endosomal carrier vesicle fuses with late endosome.
c. Ligand and receptor are transferred to lysosome for destruction.
3c4) LDLs and cholesterol metabolism
An example of receptor-mediated endocytosis delivering nutrients.
Cells need cholesterol for membrane assembly or metabolic processes (e.g. steroid
metabolism).
Cholesterol is synthesized by liver.
Hydrophobicity of cholesterol prevents transport in blood as free cholesterol.
BIOL 130
MODULE 3
LDL particles carry cholesterol in blood from liver to body's cells.
LDL = low density lipoprotein
LDL particles (Karp: p. 307 Fig. 8.43)
Consists of a central core of cholesterol esterified to long-chain fatty acids.
Core is surrounded by
a.
Coat of phospholipids and unesterified cholesterol.
b. A single molecule of apolipoprotein B.
Apolipoprotein B interacts with LDL receptors.
LDL receptors
Cells contain a variable number of LDL receptors on plasma membrane.
Concentrated in coated pits.
Once LDL binds receptors:
a.
Pit invaginates to form a coated vesicle.
b. Clathrin coat is disassembled.
c.
LDL receptors are recycled back to plasma membrane
d. LDL particles are delivered to lysosomes.
e.
Protein of LDL is digested and cholesterol is released.
3c5) LDL and HDL in heart disease
LDL and heart disease
High blood levels of LDL are associated with increased risk of heart disease.
(e.g.) atherosclerosis.
Atheroscleoris is characterized by LDL-containing plaques (atherosclerotic
plaques).
Plaques form on inner walls of blood vessels.
This causes the narrowing of major arteries and reduced blood flow.
Plaques also act as sites for formation of blood clots.
Blood clots that block coronary arteries are leading cause of myocardial infarction
(heart attack).
LDL and atherosclerotic plaque formation (Fig. 8.44)
1.
2.
3.
Initiated by:
i.
Injury to endothelial cells lining blood vessels.
ii. Reactive oxygen species (ROS) chemically alter LDL-cholesterol.
Macrophages are attracted to injured endothelium and
i.
Ingest LDL oxidized by ROS.
ii. Accumulate cholesterol-rich fatty droplets in cytoplasm.
iii. Are referred to as macrophage foam cells.
Smooth muscle cells around blood vessel are:
i.
Stimulated to proliferate by foam cells and
ii. Produce fibrous cap that bulges into arterial lumen.
50
BIOL 130
MODULE 3
Lowering blood LDL levels
Block key enzyme in cholesterol synthesis.
Enzyme is HMG CoA reductase.
Inhibitors are lovastatin and pravastatin.
Lower blood cholesterol lowers frequency of heart attacks.
High-density lipoproteins (HDLs)
Are similar to LDL but have different protein.
Have apolipoprotein A-I.
Circulate in blood as HDL particles.
Pick up excess cholesterol from plasma membrane of body's cells.
Deliver cholesterol from body's cells to liver.
Liver cells take them up by endocytosis and excrete them as part of bile.
HDLs and heart disease
High blood levels of HDL are associated with decreased risk of heart disease.
This is because HDL facilitates removal of cholesterol from blood.
Another factor is cholesteryl ester transfer protein (CETP).
CETP moves cholesterol from HDL to LDL.
Inhibiting CETP might elevate HDL and reduce coronary artery disease.
51
BIOL 130
MODULE 3
3c6) Phagocytosis 'cell eating' (Karp: p. 308; Fig. 8.45)
Uptake of particulate material and delivery to a lysosome.
Delivery done via a phagosome.
Lysosome is cell's digestive organelle. (Karp p. 297)
Phagosome fuses with primary lysosome to form secondary lysosome.
Fusion activates lysosomal enzymes.
~50 different hydrolytic enzymes in lysosome. (Karp: p. 298; Table 8.1)
Contents digested providing:
a.
Nutrition products move into cytoplasm.
b. Defense microorganisms are killed.
Macrophages and neutrophils (Karp p. 308-309)
are professional phagocytes in animals.
have mechanisms for killing ingested microorganisms:
1. lysosomes contains lysozyme, an enzyme that degrades bacterial cell walls.
2. Acidic pH of lysosome kills some.
3. oxygen free radicals generated within phagosome kill some bacteria.
a. hydrogen peroxide yields superoxide radical. (Karp p. 34)
b. nitric oxide (NO) yields peroxynitrite (Karp p. 640-641)
3c7 Some bacteria circumvent being killed by macrophages (Karp p 308-309)
1.Mycobacterium tuberculosis causes tuberculosis
- engulfed into phagosome but prevents fusion of phagosome with
lysosome.
2. Coxiella burnetii causes Q fever.
- this bacteria is not killed by either lysosomal enzymes or acidic pH.
3. Listeria monocytogenes causes meningitis
- produced proteins that destroy the integrity of the lysosomal membrane
allowing the bacteria to escape into the cytoplasm.
52
BIOL 130
MODULE 3
53
3d: ATP Formation and the Mitochondria: Unit #13
(Karp: p. 173-209)
3d1) Fermentation: Anaerobic oxidation of pyruvate (Karp: p. 111 Fig 3.29; p 177 Fig. 5.5)
1.
Glycolysis glucose to pyruvate
a.
In cytosol
b. Presence or absence of O2
c.
Net of 2 ATP/glucose
2.
Fermentation
a.
In cytosol
b. Absence of O2
c.
d.
e.
Regenerates NAD+ to support glycolysis
Muscle
i.
Lactate dehydrogenase
ii. Lactate
Yeast
i.
Alcohol dehydrogenase
ii. Ethanol
3d2) Aerobic Oxidation of Pyruvate (Fig. 5.5)
1.
2.
3.
When O2 is present, pyruvate enters mitochondria.
Completely oxidized to CO2.
a.
Pyruvate dehydrogenase
b. TCA or Krebs cycle
Get up to 36 ATP/glucose
Oxidation of pyruvic acid to Acetyl CoA
1.
2.
3.
Catalyzed by pruvate dehydrogenase complex
One CO2 is evolved.
NADH is formed
3d3) TCA or Krebs cycle (Karp: p. 179 Fig. 5.7)
1.
2.
3.
2-carbon acetyl group condenses with 4-carbon oxaloacetate to form 6-carbon citrate.
One turn of cycle evolves 2 CO2.
his completes oxidation of pyruvate.
BIOL 130
4.
MODULE 3
Free energy is conserved in.
a.
3 NADH and 1 FADH2.
Will be used in electron-transport chain to form ATP.
b. 1 GTP energetically equivalent to ATP
TCA or Krebs cycle: Central pathway of cell (Karp: p. 180 Fig. 5.8)
1.
2.
3.
Metabolites of other catabolic pathways enter TCA cycle
e.g. breakdown of proteins
TCA cycle metabolites can be used to synthesize larger molecules (anabolism).
e.g. amino acids for protein synthesis
Amphibolic pathway used in both catabolism and anabolism.
3d4) Mitochondria structure (Karp: p. 176 Fig. 5.3; Fig. 5.4)
1. Outer membrane
a.
Porins
Integral proteins that form large, nonselective membrane channels.
2. Intermembrane space
3. Inner membrane
a.
Electron transport chain
b. ATP synthase
4. Matrix
a.
TCA cycle
b. DNA (genes for ~13 polypeptides)
c.
Ribosomes
3d 5) Electron transport chain (or respiratory chain) (Karp: p. 188 Fig. 5.17)
1. Sequential transfer of electrons from one electron carrier to another.
2. All carriers are associated with the inner mitochondrial membrane.
3. Five types of electron carriers
Components of electron transport chain (Karp: p. 185 Fig. 5.12)
1. Flavoproteins
Prosthetic groups are derived from riboflavin. (FAD or FMN)
e.g. NADH dehydrogenase
2. Cytochromes
Proteins that contain heme groups.
Iron of heme undergoes reversible transition between Fe3+ and Fe2+ .
3. Ubiquinone (or coenzyme Q)
Lipid-soluble molecule
4. Iron-sulfur proteins
Iron is closely linked to inorganic sulfur.
5. copper in cytochrome oxidase
54
BIOL 130
MODULE 3
55
Function of Electron Transport Chain
1.
2.
3.
4.
Free energy released from oxidation/reductions of electron transport chain moves
H+ from matrix to intermembrane space.
This sets up a H+ gradient.
Ionic gradient across a membrane represents a form of energy.
Energy is used to form ATP. This is oxidative phosphorylation.
Proton-motive force
1.
2.
3.
4.
Two components to H+ gradient across inner mitochondrial membrane.
a.
pH gradient (chemical gradient)
b. Voltage gradient (electrical gradient)
Therefore, this is an electrochemical gradient.
Proton-motive force ( )
Expression of energy present in electrochemical gradient.
~220 mV
Voltage component ~80 %; pH component ~20 %
Maintenance requires inner mitochondrial membrane be highly impermeable to H+.
Electron transport is uncoupled from ATP formation by 2,4 dinitrophenol.
Makes inner mitochondrial membrane permeable to H+.
also drives ADP, phosphate and Ca++ into matrix.
3d 6) ATP formation
1. Chemiosmotic mechanism
Energy stored in proton gradient drives phosphorylation of ADP.
2. Catalyzed by ATP synthase.
3. ATP synthase (Karp: p. 194 Fig. 5.23)
a.
F1 headpiece projects into matrix.
b. F0 basepiece embedded in lipid bilayer contains H+ channel.
4. Controlled movement of H+ through channel induces:
a.
Conformation changes
b. These drive ATP formation.
3d7) Overall Products from oxidizing glucose
Overall products of electron transport chain
1.
2.
ATP
Each NADH = 3 ATP
Each FADH2 = 2 ATP
H2 O
Formed by O2 finally accepting electrons.
BIOL 130
MODULE 3
Return from complete oxidation of 1 molecule of glucose
Glycolysis glucose to pyruvate = 2 ATP
(also two NADH2)
Pyruvate acetyl CoA
(do twice) = 2NADH2
Two turns of Krebs cycle
6 NADH
2 FADH2 = 4 ATP
2 GTP
= 6 ATP
= 18 ATP
= 2 GTP
2NADH from glycolysis
= 4 ATP
(glycerol phosphate shuttle; Karp 181; Fig. 5.9)
Total
= 36
56
BIOL 130
MODULE 3
57
3e: Photosynthesis and the Chloroplast: Unit #14
(Karp: p. 206-223)
3e1) Heterotrophs vs. Autotrophs
Heterotrophs:
Depend on an external source of organic compounds.
Earliest life forms must have been heterotrophs.
Earliest life forms would have utilized organic molecules that had formed abiotically.
(see Fig 1.9 on p 9 Karp)
Autotrophs:
Utilize CO2 to manufacture their organic molecules.
Chemoautotrophs
Utilize the energy stored in inorganic molecules (e.g. hydrogen sulfide) to
convert CO2 into organic compounds.
Phototrophs
Utilize the radiant energy of sun to convert CO2 into organic compounds.
Photoautotrophs include
1.
2.
3.
4.
Higher plants
Eukaryotic algae
Various flagellated protists.
A variety of prokaryotes (e.g. green bacteria)
All carry out photosynthesis
Photosynthesis
Sunlight is transformed into chemical energy and utilized to form carbohydrates.
Light
CO2 + H2O
(CH2O) + O2
Photosynthesis in higher plants will be examined.
BIOL 130
MODULE 3
58
3e2) Chloroplast
1. Organelle in which photosynthesis takes place.
2. Located predominantly in mesophyll cells of leafs.
3. Semi autonomous and self replicating.
4. Bound by 2 membranes separated by a narrow space.
5. Thylakoids flattened membranous sacs within chloroplast.
a.
Lumen space inside a thylakoid
b. Grana orderly stacks of thylakoids.
6. Stroma space surrounding thylakoids.
Overview of photosynthesis
light
6CO2 + 12 H2O C6H12O6 + 6O2 + 6H2O
2 series of reactions.
1.
2.
Light reaction (light-dependent reaction)
a.
Energy from sunlight is converted to chemical energy.
b. Products.
i.
ATP
ii. NADPH
c.
Occurs in thylakoid membranes.
Dark reaction (light-independent reaction)
a.
ATP and NADPH are used to synthesize carbohydrates.
b. Occurs in stroma.
3e3) Absorption of light
Energy comes from sun in form of electromagnetic radiation.
These radiations travel in discrete packets called photons.
When a photon is absorb, compound is converted to a higher energy state (excited state).
Ground state may be re-established in three different ways:
1. Energy may be dissipated as heat.
2. Energy may be reemitted in form of longer wavelength, fluorescence.
3. Energy may be transferred to another molecule.
This is what happens with photosynthetic pigments.
Photosynthetic Pigments
Pigments are molecules that contain a chromophore.
Chromophore chemical group capable of absorbing light of particular wavelengths ().
Absorption spectrum plot of intensity of light absorbed vs. .
Action spectrum plot of physiological response vs. .
Action spectrum of photosynthesis (Fig. 6.8).
Follows absorption spectrum of the chlorophylls and carotenoids.
BIOL 130
MODULE 3
59
1.
Chlorophylls: major light capturing molecules
Absorb light of blue and red (Karp: p. 212 Fig. 6.6)
Reflects green (we see)
Two parts (Fig. 6.6):
a.
Porphyrin ring functions in light absorption
Magnesium atom in center of ring
Different side groups on ring.
This gives different kinds of chlorophyll (a b c & d)
b. Phytol side-chain
Inserts chlorophyll in lipid bilayer of thylakoid membrane.
2.
Carotenoids: an accessory pigment (Karp p. 212)
Long hydrocarbon chains containing alternating double bonds.
Increase efficiency by absorbing light in those regions where chlorophyll absorbs
light inefficiently.
Absorb light of blue and green (400-550nm) .
Reflects yellow, orange and red .
Protect photosynthetic machinery from damage caused by reactive oxygen species.
3e4) Reaction-center chlorophyll (Karp: p. 213 Fig. 6.9)
Specific chlorophylls capable of transferring electrons to an electron acceptor.
Other pigments act as a light-harvesting antenna system.
Absorb light at other and transfer energy to reaction center chlorophyll.
Reaction-center chlorophylls are organized into 2 photosystems.
These linked by a chain of electron carriers.
All within thylakoid membranes.
Photosystem II (PSII) (Karp: p. 215 Fig. 6.11 but not all details)
1.
2.
3.
4.
Reaction center chlorophyll is referred to as P680.
P = pigment
680 = of light that this chlorophyll molecule absorbs most strongly.
When P680 absorbs photons, it gives up electrons to a primary electron acceptor
(pheophytin) of higher reducing potential.
P680 replenishes its electrons from H2O.
As H2O becomes oxidized, O2 is released.
Pheophytin eventually transfers electrons to a chain of electron carriers.
BIOL 130
MODULE 3
Photosystem I (PSI) (Karp: p. 218 Fig. 6.15 but not all details)
Reaction center chlorophyll is P700.
P700 accepts electrons from last member of electron transport chain of PSII.
P700 is raised to an excited state by absorbing light.
In this state electrons are transferred to primary electron acceptor A0.
A0 has a high reducing potential.
Electrons have two fates. They can pass down:
a.
A short electron transport chain to NADP+ to form NADPH.
b. To P700 to form ATP (cyclic photophosphorylation). (Fig. 6.17)
Z scheme or pathway (Karp: p. 214 Fig. 6.10; p. 217 Fig. 6.14)
1.
2.
3.
4.
Two photosystems acting in series.
Electron flow occurs in 3 steps:
a.
Between H2O and PSII.
b. Between PSII and PSI. Electron transport chain
(Fig. 6.16 but not all details)
c.
Between PSI and NADP+.
As electrons flow along Z-pathway, H+ ions are moved from stroma to inner
compartment of thylakoids.
Important end result is proton gradient.
Proton concentration:
a.
High in lumen of thylakoid.
b. Low in stroma.
3e 5) Photophosphorylation (Karp: p. 219 Fig. 6.16)
1.
2.
3.
4.
Formation of ATP as result of electrons moving through photosystems I and II.
As in mitiochondria:
a.
Proton gradient drives ATP formation.
b. Enzyme is ATP synthase.
ATP synthase embedded in thylakoid membranes.
Two types of photophosphorylation
a
Noncyclic photophosphorylation
b. Cyclic photophosphorylation
Noncyclic photophosphorylation (Karp: p. 214 Fig 6.10; p. 219 Fig. 6.16)
1.
2.
3.
Electrons move in linear path from H2O to NADP+
Uses photosystems I and II
Formation of ATP, NADPH and O2
60
BIOL 130
MODULE 3
Cyclic photophosphorylation (Karp: p. 221 Fig. 6.17)
1. Electrons move from P700 to ferredoxin and back to P700
2. Involves photosystem I only.
3. Formation of ATP
Relative amount of noncyclic vs. cyclic photophosphorylation is regulated.
3e 6) CO2 fixation and formation of carbohydrate
Done in all photosynthetic plants by C3 cycle.
C3 or Calvin cycle (Karp: p. 222 Fig. 6.19; p. 223 Fig. 6.20)
1. Carboxylation
a.
CO2 combines with ribulose -1, 5-bisiphosphate.
b. Forms a transient 6 carbon compound.
c.
This breaks down to form 2 molecules of 3-phosphoglycerate (PGA).
d. Catalyzed by ribulose -1, 5-bisphosphate carboxylase ("Rubisco").
2. ATP is used to form 1, 3 bisphosphyglycerate .
3. NADPH is used to reduce above to glyceraldehyde 3-phosphate (GAP)
GAP has a number of fates:
1.
2.
Remain in chloroplast.
a.
Regenerate RuBP
b. Converted to starch
Exported to cytosol.
a.
Converted to sucrose.
b. Oxidized in glycolysis and TCA cycle to provide ATP
61
BIOL 130
MODULE 4
Module 4 outline and notes: Flow of Genetic Information
4a Flow of information in cells
4a1) The Gene as an Abstract Concept
4a2) Chromosomes: Physical Carriers of the Genes
4a3) How do genes determine a characteristic of an organism?
4a4) What is the chemical nature of the gene
4a5) How does DNA direct synthesis of proteins?
4a6) Central Dogma of Molecular Biology
4a7) Genetic Code
4a8) Modern attempts to define a gene
4a9) Genome
4a10) Organization of nuclear genome: chromatin and chromosomes
4a11) Biochemistry of Chromatin
4a12) Nucleosomes
4a13) Jumping genes
4a14) Reversing the normal flow of genetic information: reverse transcription
4b Transcription and translation
4b1) Basic Transcription Process
4b2) DNA sequences critical for starting and stopping transcription
4b3) Proteins responsible for transcription
4b4) Describing RNAs
4b5) RNA processing
4b6) Mature mRNAs
4b7) Translation of mRNA or protein synthesis
4b8) Ribosomes
4b9) Posttranslational modifications
4c Control of gene expression
4c1) Control of gene expression in eukaryotes
4c2) Nuclei of cells of vertebrates are totipotent
4c3) Genomic Control
4c4) Transcriptional-level Control
4c5) Glucocorticoid receptor: an example of transcriptional activation
4c6) Transcriptional repression
4c7) Processing-level control
4c8) Translational-level control
4c9) Posttranslational control
4d DNA replication and repair
4d1) Modes of DNA replication
4d2) DNA replication in bacteria
4d3) Enzymes of DNA replication
4d4) Semidiscontinuous replication
4d5) DNA replication in eukaryotic cells
4d6) Central Dogma of Molecular Biology
4d7) DNA repair
62
BIOL 130
MODULE 4
4e Cell cycle
4e1) Cell cycle defined
4e2) Macromolecular synthesis during the cell cycle
4e3) Cell Cycles in vivo
4e4) Control of cell cycle
4e5) Cyclin-dependent kinases (Cdks) and cell cycle control
4e6) Cell Cycle Check Points
4e7) M phase: Mitosis and Cytokinesis
4e8) Meiosis
63
BIOL 130
MODULE 4
64
4a: Flow of Information in Cells Unit #15
(Karp: p. 379-381; 386-390; 419-422; 455-457; 481-487)
4a 1) The Gene as an abstract concept (Karp: p. 388-390)
Mendel in 1860s.
A gene is a unit of heredity.
A gene is an element that controls a characteristic or a feature of an organism.
Mendel's laws describe behavior of these abstract units
4a 2) Chromosomes: Physical Carriers of the Genes (Karp: p. 380-381)
Chromosomes (cytologist's definition)
Organization of nucleus into visible threads at time of cell division (colored bodies).
Humans have 23 (haploid) distinct pairs for a total of 46 chromosomes (diploid)
Genes are carried by chromosomes Sutton in 1900
1.
2.
3.
Like genes chromosomes come in pairs.
Paired or homologous chromosomes separate during gamete formation.
Like genes chromosomes are carefully replicated and separated during cell division.
4a 3) How do genes determine a characteristic of an organism?
Genes function by directing the synthesis of proteins. (Karp: p. 390 Fig 10. 11)
Beadle and Tatum 1940
One gene one enzyme hypothesis
4a 4) What is the chemical nature of the gene? (Karp: pp. 386-390)
Avery, Macleod and McCarty in 1944 showed genetic material is DNA.
Accepted in 1950s when double-helix structure of DNA was determined.
4a 5) How does DNA direct synthesis of proteins?
Information stored in a gene is transferred to a mRNA.
Base or nucleotide sequence of a mRNA is complementary to its gene.
mRNA directs placement of amino acids into a protein.
Amino acid placement requires ribosomes and tRNA.
Ribosomes are complexes of rRNA and proteins.
tRNA helps translate mRNA
BIOL 130
MODULE 4
65
4a 6) Central dogma of molecular biology (Karp: p. 390; 421 Figs. 10.11; 11.2)
DNA
Transcription
mRNA
Translation
Protein
4a 7) Genetic code
Nucleotide combinations that specify placement of amino acids.
How many bases or nucleotides specify an amino acid?
1. If one base = one amino acid
2. If two bases = one amino acid we would have 16 words
42 = 16 code words are possible.
3. If three bases = one amino acid
43 = 64 code words are possible
Therefore, three bases = codon or code word.
Assigning code words
What amino acid (aa) does each code word specify?
Artificial or synthetic mRNA or homopolymer experiment
tRNA + ribosomes + aa + uuuuuu polyphenylalanine
Therefore code word for phenylalanine is uuu
Genetic code summary (Karp: p. 457; Fig. 11.41)
1.
2.
3.
4.
The genetic code is a nonoverlapping sequence of nucleotide triplets.
(overlapping code occurs in some viruses).
Genetic code is degenerate or redundant.
UGA, UAA, and UAG are chain terminators or stop codons.
Genetic code words appear to be universal.
4a 8) Modern attempts to define a gene
1. DNA segment (sequence) that carries the information for:
A single polypeptide or
A single rRNA molecule or
A single tRNA molecule
2. Entire DNA sequence that is necessary for synthesis of polypeptide or a functional
RNA sequence.
BIOL 130
MODULE 4
4a 9) Genome (Karp, p 393)
- total genetic information carried by a cell or an organism.
- genome size is measured in nucleotide (or base) pairs
1.nuclear genome
-nucleotide pairs per haploid set of chromosomes
Human nuclear genome contains:
~ 3 x 109 nucleotide pairs
~ 25,000 protein-coding genes (~1.5 % of total DNA)
2. organelle genomes- total genetic information in an organelle.
a. chloroplast genome (plants only)
~ 120,000 nucleotide pairs and 60-200 genes.
b. mitochondrial genome (plants & animals)
~ 17,000 nucleotide pairs and 37 genes. In humans.
4a 10) Organization of nuclear genome: chromatin & chromosomes
(Karp: pp. 481-487)
Nuclear genetic material of eukaryotic cells exists in two states:
1.
Interphase nucleus
Interphase is period from division to division (cell cycle).
Exists for ~23 h in proliferating mammalian cells.
DNA of interphase nucleus is complexed with proteins.
This complex is called chromatin.
2.
Chromosomes
Appear only at time of cell division.
Appear for ~1 h in mammalian cells.
Consist of a more complex organization of chromatin.
Microscopic view of chromatin
1.
Fine fibers distributed throughout interphase nucleus.
a.
Euchromatin chromatin that remains in open networks.
b. Heterochromatin
Chromatin that remains tightly condensed during interphase.
Genetically inactive.
i.
Constitutive heterochromatin
Remains in condensed state in all cells at all times.
Situated in and around centromere.
ii. Faculative heterochromatin
Specifically inactivated during certain developmental phases.
(Karp: p. 486; Fig. 12.13). e.g. X chromosome
66
BIOL 130
MODULE 4
67
4a 11) Biochemistry of chromatin
1.
2.
3.
DNA
Histones (Table 12.1, Karp: p. 481)
Highly conserved, basic proteins.
H1, H2A, H2B, H3, H4
Nonhistone chromatin proteins.
Includes transcription factors
4a 12) Nucleosomes (Karp: p. 482; Fig. 12.8; p. 485 Fig. 12.12)
1.
Subunit structure of chromatin.
2.
A nucleosome consists of:
a.
Core particle:
i. two molecules each of H2A, H2B, H3 & H4.
ii. 146 base pairs of DNA wrapped around core.
b. Linker DNA (40-60 base pairs) with linker histone (H1)
3. Nucleosomes package DNA
Packing ratio = length of DNA molecule/length of DNA in fiber
Nucleosome gives a packing ratio of ~7
Solenoid or 30 nm filament (Karp: p. 484 Fig 12.10; p. 485 Fig. 12.12)
Coiling of nucleosomal filament into higher-order, thicker filament.
Stabilized by interactions of centrally located H1.
40 fold greater packing than DNA
Chromosomes (Karp: p. 485 Fig. 12.12)
A much tighter packing of chromatin that is achieved just prior to cell division.
Packing ratio in range of 15,000 to 20,000
4a13 Jumping genes (Karp p; 402-403)
1. transposable elements (transposons)
- a segment (sequence) of DNA that can move from one location in a chromosome
to anther location either in the same chromosome or into another chromosome.
(Fig 10.26)
- can be defined by sequences
- ends contain a sequence that is inversely repeated.
2. human genome
2 main families of transposable sequences
a. DNA transposons ( Karp p 403; Fig. 10.26)
"cut and paste"
b. retrotransposons
"copy and paste" of transposons
BIOL 130
MODULE 4
68
99 % do NOT move
However, a particular form of hemophila is caused by the insertion of a
retrotransposon into the gene that encodes a key blood-clotting enzyme.
4a14 Reversing the normal flow of genetic information: reverse transcription (Karp
p; 402-404)
1. defining step is use RNA as a template to synthesize DNA.
2. enzyme that carries out this is reverse transcriptase or RNA- dependent DNA
polymerase.
3. reverse transcription occurs in nature.
a. a small amount in animal cells
i. telomerase - an enzyme that maintains telomeres
-telomeres are repeated sequences that form a cap at each end of a
chromosome (Fig. 12.20 & unit 18 Karp 494).
- is a reverse transcription that has its own small amount of RNA.
ii. a few retrotransposons code for their own reverse transcriptase.
(Fig. 10.26)
b. retroviruses
BIOL 130
MODULE 4
69
4b):Transcription and Translation: Unit #16
(Karp: p. 422-455; 461-468)
4b 1) Basic Transcription Process (Karp: p. 424; Fig. 11.4)
1.
2.
3.
4.
5.
DNA sequence of bases is copied into a RNA sequence of bases.
A
T
C
G
C
G
T
A
A
T
G
C
Enzymes responsible are DNA-dependent RNA polymerases or simply RNA
polymerases.
They utilize ribonucleotide triphosphates and require a DNA template.
Polymerases move in 3' to 5' direction along template DNA strand.
RNA grows in 5' to 3' direction.
4b 2) DNA sequences critical for starting and stopping transcription
Promoter of a gene (Fig. 11.7)
1.
2.
3.
4.
5.
A DNA sequence to which RNA polymerase binds prior to initiating transcription.
Contains information that determines which of two strands is transcribed.
Contains information that determines site at which transcription begins.
Regulates rate of gene expression.
Contains consensus sequences.
Consensus sequences are most common versions of a DNA base sequence
occurring in slightly different forms at different locations.
Location of a promoter (Figs, 11.6, 11.7, 11.18)
1.
2.
3.
4.
Nucleotide at which transcription is initiated is + 1.
To the right of +1 is downstream. (indicated by +)
To the left of +1 is upstream. (indicated by -)
Promoters usually are upstream.
Termination region
A nucleotide sequence that dictates the termination of transcription.
BIOL 130
MODULE 4
70
4b 3) Proteins responsible for transcription
RNA polymerases make RNA
1 type of RNA polymerase in prokaryotes
3 distinct types of RNA polymerases in eukaryotes (see Table 11.1, p. 426)
1. RNA polymerase I
Makes large (28S, 18S and 5.8S) ribosomal RNAs.
2. RNA polymerase II
Makes mRNAs
Makes most small nuclear RNAs (snRNAs & snoRNAs) & microRNAs.
3. RNA polymerase III
Makes small RNAs, including tRNA and 5S ribosomal RNA.
Transcription factors (Fig. 11.18)
Auxiliary proteins that assist RNA polymerases.
These bind to promoters.
General transcription factors
Required for polymerase to initiate transcription.
Specific transcription factors
Determine rate at which a particular gene or group of genes is transcribed.
4b 4) Describing RNAs
1.
Size of RNA and ribosomes described by their sedimentation coefficient.
Sedimentation coefficient is a measure of how rapidly they sediment in an
ultracentrifuge.
This is expressed in Svedberg units (S).
S units depend on both size and shape and are not arithmetic.
e.g. eukaryotic ribosome is 80S and made up of a large (60S) and small (40S)
subunit.
2.
Classic types of RNA involved in protein synthesis
Ribosmal RNA or rRNA
Messenger RNA or mRNA
Transfer RNA or tRNA
3.
Small nuclear RNAs involved in RNA processing
snRNAs (snurps) are involved in processing mRNA.
snoRNAs are involved in processing of pre-rRNA.
4.
micro RNA (miRNAs) a newly discovered network for gene regulation
- roughly 20-23 nucleotides in length of dsRNA.
- may be 1000s of distinct miRNAs.
BIOL 130
5.
MODULE 4
71
small interfering RNAs (siRNAs)
- 21-23 nucleotide segments of double stranded (ds) RNA.
- derived from double stranded product of:
a. an RNA virus
b. transposable element (mobile genetic elements, such as retrotransposons)
c. dsRNA by a researcher
common features of miRNAs and siRNAs (cousins)
both act to silence the expression of cytoplasmic mRNA
piwi (piRNAs) are a new slightly longer class of siRNAs
piRNA suppress transposable elements in the germ line
6. Other noncoding RNAs (ncRNAs) (Karp: p 454-455)
- new research area.
- might have diverse regulatory activities.
4b 5) RNA processing
RNAs are usually shortened and chemically modified after synthesis and before
use.
This usually occurs after transcription and is described as posttranscriptional processing.
Initial RNA is called primary transcript.
Final RNA is sometimes called mature RNA.
Many primary transcripts can be divided into two types of sequences:
1. Introns will be removed from primary transcript and often destroyed
2. Exons will be spliced together to form mature RNA
RNA processing occurs in tRNA, rRNA, and mRNA.
Will study mRNA (Karp p. 443 Fig 11.29)
Capping or 5' cap of mRNA (Karp: p. 437 Fig. 11.21)
All eukaryotic mRNAs have methylguanosine at 5' end.
Consists of a methylated quanylate stuck onto the chain backwards
Roles of 5' cap:
a.
Transporting mRNA out of nucleus
b. In initiating mRNA translation
Poly A tail of mRNA (Fig. 11.29)
Most but not all eukaryotic mRNAs have poly A tails.
Nuclease cleaves off a portion of 3' end.
Poly(A) polymerase adds poly A to 3' end but does not use DNA template.
5'GUGAACUUU3'AAAAAAAAAA etc.
Tail is up to 200 nucleotides long
Poly A tail provides stability by protecting mRNA from premature degradation.
BIOL 130
MODULE 4
72
Split genes (Figs. 11.23, 11.29)
Genes for many eukaryotic mRNA are split.
Genes contain intervening sequences.
Intervening sequences or Introns are regions of DNA not expressed in mature mRNA.
Exons are regions of DNA expressed in mature mRNA.
mRNA splicing (Figs. 11.29, 11.35)
Initial or primary mRNA transcripts are often very large, have diverse base
sequence, and found only in nucleus.
These are heterogeneous nuclear RNAs (hnRNAs).
HnRNAs are precursors to mRNA.
Splicesosomes act on hnRNAs.
Splicesosomes contain snRNAs and proteins (ribonucleoprotein particles or snRNPs)
Some snRNAs are ribozmes.
Ribozmes are RNA catalysts that act on RNA to cut up RNA.
Splicesosomes cut and splice hnRNAs to yield final mRNAs.
BIOL 130
MODULE 4
73
4b 6) Mature mRNAs (Karp: p. 437 Fig. 11.21; p. 443 Fig. 11.29)
Contain continuous sequence of nucleotides encoding a specific polypeptide.
Contain a significant noncoding segment.
Noncoding portions are found at both 5' and 3' ends
Noncoding sequences are called untranslated regions (UTRs)
UTRs have regulatory roles.
4b 7) Translation of mRNA or protein synthesis (Fig. 11.46)
Polymerization of proper sequence of amino acids.
Translation starts at AUG and continues to 1 of 3 possible stop codons.
Involves tRNAs and ribosome.
Ribosome moves down and reads mRNA 1 codon at a time.
Polypeptide grows out from ribosome.
Grows N terminus to C terminus.
tRNA (Karp: p. 458 Fig. 11.42; p. 459 Fig. 11.43)
Their function is to recognize a mRNA codon and to position correct amino acid
into growing polypeptide.
Two important sites:
1. Amino acid attachment site. at 3' end.
Attachment of amino acid makes tRNA an aminoacyl (aa)-tRNA.
2. Anticodon
aa-tRNA recognizes correct mRNA codon during translation through
pairing between 3 complementary bases in mRNA codon and tRNA
anticodon. (Karp: p. 461 Fig. 11.46)
4b 8) Ribosomes (Karp: p. 77 Fig. 2.56; p. 428; Fig. 11.9)
rRNA protein complexes.
Ribosomes are made up of two different subunits.
In eukaryotes these are 60s and 40s.
Together form a functional unit, which is 80s.
Eukaryotic ribosomes have 4 distinct rRNAs.
a.
Large ribosomal subunit has 3.
28S
5.8S
5S
b. Small ribosomal subunit has 1: 18S
The gene for 28S, 18S & 5.8S is in nucleolus.
5S rRNA genes are located outside nucleolus.
Formation of polyribosomes or polysomes (Karp: p. 421; Fig. 11.2; p. 467; Fig. 11.50)
A complex of mRNA and many ribosomes.
BIOL 130
MODULE 4
mRNA being read by several ribosomes simultaneously.
Thus several polypeptide chains can be made from it simultaneously.
Greatly increases rate of protein synthesis.
4b 9) Posttranslational modifications
1.
2.
3.
4.
Polypeptide is often cleaved.
Addition of groups (ex phosphate) to R group of amino acid residues.
Polypeptide becomes properly folded.
Folding can occurs spontaneously but is assisted by chaperones and protein
disulfide isomerases.
74
BIOL 130
MODULE 4
75
4c: Control of Gene Expression: Unit #17
(Karp: pp. 503-531; 448-451)
4c 1) Control of gene expression in eukaryotes
Typical mammal has protein-coding information for as many as 25,000 polypeptides.
Typical mammal has ~250 different cell types.
Typical mammalian cell makes ~5,000 different polypeptides at any given time.
Different cell types will make slightly different sets of polypeptides.
Yet nuclei of different cell types have same information.
4c 2) Nuclei of cells of vertebrates are totipotent.
Experimental proof.
A somatic cell nucleus from a frog supports development of a frog if placed in the
appropriate cytoplasm cytoplasm of an unfertilized egg.
A similar experiment has now been done with mammals (Karp: p. 504, Fig. 12.31).
How is it that different types of cells in an animal or plant are able to synthesize
different proteins?
4c 3) Genomic Control (relatively rare)
1.
2.
3.
Loss of DNA (never occurs in mammals)
Selective amplification of genes.
ex. genes for rRNA are amplified in oocyte during oogenesis.
Rearrangement of DNA sequences.
This occurs in antibody formation.
4c 4) Transcriptional-level control (Karp: p. 505, Fig 12.32)
single most important mechanism.
Differential gene transcription: genes are turned either on or off.
transcriptional activation turning genes on.
involves interaction between DNA sites and proteins.
DNA sites involved in regulating transcription (cis-acting elements)
1. Promoters (Fig 12.40)
a. DNA sequences to which RNA polymerase II binds prior to initiating
transcription.
b. found upstream of a gene.
c. contain promoter elements smaller sequences that are required for full
promoter activity.
e.g. TATA box
d. promoter divided into segments
e.g. core promoter TATA box to transcription start site.
2.
Enhancers
BIOL 130
MODULE 4
76
a. DNA sequences that provide specificity of time, place and level of
expression of a specific gene.
b. Stimulate transcription above basal level.
c. Can function at variable distances from start of transcription.
Transcription factors (Trans-acting elements)
Proteins that modulate transcription
Two functional classes:
a.
General transcriptional factors:
Bind at core promoter sites in association with RNA polymerase.
b. Specific transcription factors
Either stimulate or inhibit transcription of specific genes.
Bind at specific DNA sites- response elements
Two portions or domains:
a.
Transcription activation domain portion that is required for activation of
transcription.
b. DNA binding domain portion that recognizes and interacts with DNA
consensus binding site.
Any one of several structural motifs. zinc finger and helix-loop-helix (Karp:
p. 510-511 Figs. 12.37; 12.38)
4c 5) Glucocorticoid receptor: an example of transcriptional activation
1. glucocorticoids (stress hormones)
a. steroids made by adrenal gland.
b. cortisol is one.
c. act through glucocorticoid receptor (GR)
d. GR is a transcription factor.
e. GR binds glucorticoid response element (GRE).
2. Phosphoenolpyruvate carboxykinase (PEPCK)
a. is key enzyme in gluconeogenesis.
b. Gluconeogenesis converts pyruvate to glucose. (Karp p 114, Fig 3.31)
c. PEPCK levels elevated by certain hormones (e.g. glucocorticoids).
3. Regulatory DNA sequences for PEPCK (Karp p 515, Fig. 12.43)
a. Core promoter elements (found in many genes)
TATA box Site of assembly of general transcription factors.
CAAT box
GC box
b. Distal promoter elements
a.
Glucocorticoid receptor glucocorticoid response element (GRE)
b. Thyroid receptor
c.
Insulin
4. Regulation of expression by glucocorticoids (Karp: p. 515 Fig. 12.43)
a. Cortisol (a lipid) diffuses through plasma membrane and binds GR.
b. Upon binding cortisol, GR is activated, moves to nucleus and binds GREs.
c. This increases rate expression of PEPCK.
d. promotes conversion of amino acids to glucose.
BIOL 130
MODULE 4
77
4c 6) Transcriptional repression (Karp: p. 498-499; 528-529)
the shutting off of genes.
usually groups of genes
not well understood
e.g. X chromosomes of female mammals.
1. repression through chromatin changes
a. Chromatin modifications
i. histones undergo several types of posttranslational modifications.
e.g. acetylation, methylation, phosphorylation
b. Deacetylation -removal of acetyl groups
i. deacetylation of H3 and H4 is initial step in conversion of euchromatin to
heterochromatin (genetically inactive). (Karp p 488; Fig. 12.16)
ii. done by histone deacetylases (HDACs)
c. methylation of H3
i. occurs on lysine at position 9 (K9)
ii. done by histone methyltransferase
iii. accompanies deacetylation
d. chromatin can now be packaged into higher order, more compact structure.
2. DNA methylation (Karp p 529, Fig. 12.49)
a. methyl groups are attached to carbon 5 of specific cytosines in DNA of animals.
b. methylation is thought to maintain genes in an inactive state.
c. Remarkable shifts in DNA methylation occurs during the life of a mammal.
i. Zygote is substantially methylated
ii. During cleavage, genome undergoes global demethylation.
iii. After implantation, DNA is methylated again except for primordial germ cells.
iv. Somatic cells maintain the methylation pattern for life
v. Primordial germ cells give rise to sperm and egg which become methylated.
4c 7) Processing-level control (Karp: p. 531)
Same gene can be spliced differently in different cell types.
Alternative splicing generates protein diversity
e.g. gene for fibronectin (Karp: p. 531 Fig. 12.53)
Fibroblasts secrete fibronectin into extracellular matrix.
Liver cells secrete fibronectin into plasma.
BIOL 130
MODULE 4
78
4c 8) Translational-level control (Karp: p. 532-536)
movement, inhibition or destruction of mRNA.
1.
Cytoplasmic localization of mRNAs
Mechanisms often involve untranslated regions (UTRs) of mRNA.
5'UTR extends from cap to start codon, AUG.
3'UTR extends from stop condon to end of poly A tail.
3'UTR region localizes mRNAs in Drosophila egg.
This contributes anterior-posterior axis of Drosophila. (Fig. 12.53)
mRNA for biscoid gene localized in anterior end of oocyte.
Protein for biscoid gene plays a critical role in development of head
2.
Control of mRNA translation (translation blocks)
a. masked mRNAs by proteins
Some well documented cases, e.g. sea urchin egg.
Certain proteins prevent mRNA from binding to ribosomes until after fertilization
b. microRNA (miRNA) (Karp 448-451; Fig 11.38 right hand side)
primary transcript of miRNA (pre-miRNA) fold backs on itself and is cleaved to
give double stranded miRNA.
associates with miRISC protein complex and is unwound into single strands.
this mature miRNA binds to a complementary region on a mRNA and inhibits
the translation of the mRNA.
3.
Control of mRNA stability
How long does a mRNA persist?
Longer a mRNA persists, more times it can be translated.
In eukaryotes different mRNAs have different half-lives.
e.g. from 10 mins. to 24 h.
a. One method of controlling mRNA degradation involves poly A tails.
Nibbled away by polyA ribonuclease.
When down to 30 residues, mRNA rapidly degraded from 5' end. (Karp: p. 526
Fig. 12.56)
b. RNA interference (RNAi) by siRNA (Karp 448-451; Fig 11.38 left hand side)
siRNA targets the same transcripts from which it arose.
Dicer, a particular type of ribonuclease, acts on dsRNA (e.g. RNA virus) to form
siRNAs.
siRNA associate with a protein complex (RISC) and a helicase unwinds the 2 RNA
strands.
The active, single-stranded siRNA associated with proteins of the RISC complex
binds to a mRNA target that has a complementary sequence.
the mRNA is now degraded.
BIOL 130
MODULE 4
4c 9) Posttranslational control
1.
How long does a polypeptide persist?
Varies from few minutes to weeks.
2.
Factors determining protein stability
N-end rule (Karp: p. 529-530)
Polypeptides with arginine or lysine at N-terminus are short lived
3.
Proteasomes a mechanism of degradation
Barrel-shaped, protein-degrading machines ( Karp p 529; Fig 12.58).
Consists of four rings of polypeptide subunits.
Two central rings function as proteolytic enzymes.
Targeting proteins for destruction
Small protein called ubiquitin is added to a lysine residue.
79
BIOL 130
MODULE 4
80
4d: DNA Replication and Repair: Unit #18
(Karp: pp. 533-548; 552-554)
4d 1) Modes of DNA replication
Watson and Crick suggested a semi-conservative mode. (Karp: p. 534 Fig. 13.1)
T
G
T
C
C
A
C
A
G
G
However, 3 modes of DNA replication were possible. (Karp: p. 534 Fig. 13.1;
Fig. 13.2)
1. Conservative
2. Semi-conservative
3. Dispersive or random
old/old
new/new
old/new
new/old
Semi-conservative mode proved to be correct.
4d 2) DNA replication in bacteria
1.
2.
3.
4.
Bacterial chromosome is circular DNA molecule.
Origin of replication a specific site (~245 base pairs) on DNA at which replication
begins.
Replication proceeds outward from origin in both directions, i.e. bidirectionally.
(Karp: p. 537 Fig. 13.5)
Replication forks sites where parental double helix is undergoing strand separation
and nucleotides are being added into newly synthesized complementary strands.
4d 3) Enzymes of DNA replication
1.
2.
3.
4.
DNA polymerases
a.
DNA polymerase I (primer removal and fills in gaps)
b. DNA polymerase II (unknown)
c.
DNA polymerase III (replication)
RNA polymerases
a.
general one
b. specific one called a primase
DNA ligase
Topoisomerases
Change supercoiled state of a DNA duplex.
BIOL 130
MODULE 4
81
Characteristics of DNA polymerases
Require presence of:
1. 4 deoxyribonucleotides: dTTP, dATP, dCTP and dGTP.
2. DNA template (Karp: p. 539 Fig. 13.7)
3. Primer strand with 3'OH terminus
Surprising feature of DNA polymerases
Only capable of synthesizing DNA in a 5' to 3' direction.
4d 4) Semidiscontinuous replication (Fig. 13.8; Fig. 13.9)
1.
2.
3.
One DNA strand grows toward the replication fork.
This is called the leading strand.
This is synthesized continuously.
The other DNA strand grows away from the replication fork.
This is called the lagging strand.
This strand is synthesized discontinuously as fragments.
These are Okazaki fragments.
As required by the DNA polymerase, both strands are assembled in a 5' to 3' direction.
Short, transient RNA primers (Karp: p. 541 Fig. 13.11)
Required for initiation of elongation.
For the leading strand, RNA polymerase makes them.
For Okazaki fragments, primase makes them.
After DNA elongation, RNA primers are removed and gaps filled in by DNA
polymerase I.
The replication fork (Karp: p. 536-537; Figs. 13.5, 13.6)
Separation of strands of a double helix unwinds the structure.
Because the DNA molecule is not free to be entirely rotated, DNA in rear of
replication fork becomes positively supercoiled.
DNA gyrase (a topoisomerase) relieves the tension that builds ups during
replication.
Target for antibiotic and anticancer drugs.
BIOL 130
MODULE 4
82
4d 5) DNA replication in eukaryotic cells
two significant differences from prokaryotes
1. replication occurs simultaneously at many sites along the chromosomal DNA.
(Karp: p. 547 Fig. 13.19)
These units of replication are termed replicons.
Replicons are generally 50 to 300 kilobases in length.
Each replicon has its own origin of replication.
2. end replication problem (Karp p 494; Fig 12.20)
arises because the chromosomes are linear and because lagging strand needs an
RNA primer from which to replicate back from.
When the replication fork approaches the end of a chromosome, there is no place to
lay down the RNA primer to start the very last Okazaki fragment.
problem solved by having at chromosome ends a special nucleotide sequence,
telomere.
Telomeres attract telomerase, which adds additional telomere DNA sequence to the
ends of the chromosomes.
An RNA primer is then laid down that allows the lagging strand to be completed.
4d 6) Central Dogma of Molecular Biology
Central Dogma of molecular biology
DNA
RNA
Protein
4d 7) DNA repair (Karp: p 552-553)
Cells have tremendous potential for repair.
DNA is susceptible to environmental damage
Examples
1.
2.
3.
4.
UV radiation causes pyrimidine dimers (Karp: p. 552 Fig. 13.25)
Usually thymine dimers
Ionizing radiation can break sugar/phosphate backbone.
a.
Single strand breaks
b. Double strand breaks
Free radicals convert guanine to 8 hydroxyguanine
Thermal energy of warm-blooded animals is sufficient to split adenine and guanine
bases from sugar/phosphate backbone.
Consequences if DNA lesions are unrepaired
1.
Permanent alterations or mutations in DNA
BIOL 130
2.
MODULE 4
83
a.
If in gametes, next generation is affected.
b. If in somatic cells, cells can potentially become cancerous.
Impair transcription and replication and cause cell death
Nucleotide Excision Repair
A repair mechanism for bulky lesions such as pyrimidine dimers
Mechanism: (Karp: p. 553 Fig. 13.26)
1. Pair of endonucleases make incisions in backbone of altered strand on each side of
the lesion (pyrimidine dimer).
2. DNA helicase removes damaged region away from the intact complementary
strand.
3. Gap is filled in by a DNA polymerase
4. DNA ligase joins the strands
BIOL 130
MODULE 4
4e: Cell Cycle: Unit # 19
(Karp: p. 560-571; 585-588; 590-591)
4e 1) Cell cycle defined
The time from division to division is the cell cycle.
The cell cycle is the fundamental unit of time at the cellular level
Interphase is the period from division to the next division.
What goes on during interphase?
All the components of the cell essentially double.
The cell grows.
Growth is an increase in dry mass.
Dry mass is the weight of lipids, polysaccharides, protein and nucleic acids.
Dry mass increases in an exponential pattern.
4e 2) Macromolecular synthesis during the cell cycle
1.
2.
3.
Protein synthesis
Continuous throughout the cell cycle
Individual proteins might be synthesized at a specific stage.
e.g. histones and cyclins.
RNA synthesis
Continuous except for mitosis
DNA synthesis (Karp: p. 562 Fig. 14-2)
Discontinuous there is a discrete period when DNA is replicated.
This is termed S phase.
Cell cycle stages (Karp: p. 561 Fig. 14-1)
Routine stages or phases
G1 gap 1
S phase
G2 gap 2
M mitosis (and cytokinesis)
Resting stage
G0 a block in G1
84
BIOL 130
MODULE 4
85
4e 3) Cell cycles in vivo
Cell in humans can be grouped into 3 broad categories with respect to cell cycle.
1. Cells with extreme structural specialization or differentiation.
e.g. neurons
No cycling. Instead cells are arrested in G0
2. Cells that normally do not divide but can be induced to do so.
e.g. liver cells
3
Cells that normally are cycling.
e.g. epithelial cells that line body cavities and body surface.
4e 4) Control of cell cycle
Enormous practical implications in combating cancer.
Two events are critical
1. Initiation of DNA replication
i.e. G1/S transition
2. Initiation of mitosis
i.e. G2/M transition
4e 5) Cyclin-dependent kinases (Cdks) and cell cycle control
Cyclins are regulatory subunits of specific protein kinases.
Cyclin levels vary with cell cycle stage.
High levels stimulate protein kinase activity.
(e.g. Maturation-Promoting Factor MPF in Fig. 14.4)
Protein kinase phosphorylates serine and threonine residues on proteins whose
actions are necessary for cell cycle to progress.
Slight variations of this system are found in all eurkaryotic cells.
Cyclin-dependent kinases (Cdks) are engines that drive cell cycle (see Regulation of
Cdks).
Yeast cell cycle (Karp: p. 564 Fig. 14.5)
First transition (near end of G1) is called START.
Passage through START requires activation of yeast Cdk kinase (cdc2) by one or more
G1 cyclins.
Second transition is near end of G2.
Passage through this transition requires activation of cdc2 by different group of
cyclins (mitotic cyclins).
BIOL 130
MODULE 4
Regulation of Cdks
1.
2.
3.
4.
5.
Cyclin concentration
Cdk phosphorylation state (Fig. 14.6)
Progression through cell cycle requires phosphorylation and
dephosphorylation of critical residues in cdc2 kinase.
Cdk inhibitors
Yeast a protein called Sic1
Mammalian cells one example is protein 21
Controlled proteolysis
Done by ubiquitin-proteasome pathway.
subcellular localization of cyclin
(shuttling back & forth between nucleus & cytoplasm)
4e 6) Cell cycle check points
Checkpoints are mechanisms that halt the progress of the cell cycle if:
1. DNA is damaged
2. certain critical processes have not been properly completed, such as DNA
replication during S phase or chromosome alignment during M phase.
Most proteins of checkpoint machinery have no role in normal cell cycle and are only
called into action when an abnormality appears.
Checkpoint control requires 3 distinct classes of proteins:
1. Sensors that detect abnormalities and emit an appropriate signal.
2. Transmitters that send signal along proper pathways within the cell.
3. Effectors that respond to signal and inhibit cell cycle machinery.
Example of a DNA-damage checkpoint (Karp p 568; Fig. 14.9)
Sensor: Ataxia-telangiectasia protein kinase (ATM).
1. ATM recognizes double strand DNA breaks.
2. ATM phosphorylates and activates a checkpoint kinase, Chk2.
Transmitters: Chk2 and p53
1. Chk2 phosphorylates p53 (guardian of genome).
2. Phosphorylated p53 is an active transcription factor.
3. stimulates transcription of p21
Effector:
1
p21 inhibits G1 Cdk (cyclin dependent kinase).
3. Cells are blocked in G1.
Delay in cell cycle progression:
1. Allows cell to repair and finish DNA replication.
2. Or triggers cell death if DNA is damaged beyond repair.
Checkpoints ensure that cell cycle events occur accurately and in order.
86
BIOL 130
MODULE 4
Failure of checkpoints
1.
2.
Progeny become genetically unstable.
(accumulate more and more damage).
And eventually become cancerous cells
(most cancer cells contain a number of defective genes)
(see Unit #23)
4e 7) M phase: Mitosis and Cytokinesis
Mitosis or karyokinesis nuclear division (Fig. 14.11 just for information)
Cytokinesis cytoplasmic division (Karp: p. 585-588)
Explained by contractile ring theory (Fig. 14.35)
A thin band (cortex) of contractile cytoplasm lies beneath the plasma membrane.
Cortex contains actin-myosin II filaments that interact to generate the force for
cytokinesis.
Together, mitosis and cytokinesis provide a way of transmitting information
unchanged and undiminished from mother to daughter cells.
4e 8) Meiosis reduction in chromosome number is accomplished.
(Karp Figs. 14.39, 14.4).
1.
2.
3.
Leads to formation of gametes (eggs and sperm).
Each gamete has the haploid number of chromosomes.
Not all the gametes are genetically identical. Meiosis is a source of variability.
This is why you are unique and wonderful no matter how you do in cell biology.
87
BIOL 130
MODULE 5
Module 5 outline and notes: Signal Transduction pathways
5a Cell signaling and cAMP
5a1) Cell Signaling defined
5a2) Signal Transduction pathways or reaction cascades
5a3) Two alternative types of signal transduction systems
5a4) Interaction of heteromeric G protein with receptor and effector
5a5) cAMP
5b Other 2nd messengers: lipids, calcium and nitric oxide
5b1) Lipid-derived 2nd messengers
5b2) Calcium ions as 2nd messengers
5b3) Nitric oxide (NO) as an intercellular messenger
5c Receptor Tyrosine Kinases and Cell Proliferation and Death
5c1) Receptor Tyrosine Kinases (RTKs)
5c2) EGF Receptor as a RTK
5c3) Crosstalk between Signaling Pathways
5c4) Pathways that lead to cell death (apoptosis)
5c5) External stimuli for activation phase of apoptosis
5c6) Internal stimuli for activation phase of apoptosis
5c7) Execution phase of apoptosis
5c8) Anti-apoptotic mechanisms
88
BIOL 130
MODULE 5
89
5a: Cell Signaling and cAMP: Unit #20
(Karp: p. 605-614; 618-620)
5a 1) Cell Signaling defined
Relaying of information from external environment of cells to elicit a response inside
cells.
4 very basic signaling systems:
1.
2.
3.
4.
Specific receptors in plasma membrane recognize stimulus at outer surface and
transfer signal across plasma membrane to cytoplasm (most common).
Specific receptors in plasma membrane but do not initiate signal inside the cell.
e.g. acetylcholine opens an ion channel that lies within receptor itself
(will not discuss further).
Specific receptors in cytoplasm
e.g. steroid receptors
(glucocorticoid receptor was covered. Fig. 12.43)
Intercellular messenger passes through plasma membrane.
e.g. nitric oxide (NO) (deal with in unit 21)
Basics of signaling via plasma membrane receptor
1.
2.
3.
4.
5.
Ligand agent that binds to receptor at outer cell surface.
Receptors cell surface proteins that specifically bind ligands.
Effectors bring about effects.
Second messengers substances released into the interior of the cell as the result of
the binding of a first messenger a ligand or hormone.
Signal transduction pathways or reaction cascades
Consist of a series of distinct proteins that sequentially activate or inactive
each other.
All that is transmitted across the membrane is a signal that the stimulus
(ligand) has been received.
Ligands and receptors
Ligand
a.
Extracellular ligand molecules secreted by cells
i.
Hormones
(e.g. epinephrine and glucagon).
ii. Polypeptide growth factors
(epidermal growth factor -EGF or platelet derived growth factor -PDGF)
b. Cell-to-cell contact.
c.
Extracellular matrix.
BIOL 130
MODULE 5
Receptors
Some examples
a.
Epinephrine receptor
b. EGF receptor
Effectors and 2nd messengers
Effectors
Bring about effects.
Usually an enzyme (e.g. adenylyl cyclase or phospholipase C)
Proteins with SH2 domain (a cell-signaling domain).
2nd messengers
Substances released into the interior of the cell
Often a product (cAMP) of an effector (adenylyl cyclase)
Trigger signal transduction pathways or cascades
5a 2) Signal transduction pathways or reaction cascades
Greatly amplify the original message.
Two basic types of 'on' and 'off' systems (Fig. 15.2)
1. Protein kinases/protein phosphatases (see section below)
Phosphate group either put on or taken off of proteins
2. GTP-binding proteins (or G proteins) (see next section below)
Bind either GTP (active) or GDP (inactive)
Protein kinases/protein phosphatases (Fig. 15.3)
Protein kinases
Add phosphate group to proteins.
one third of proteins are subject to phosphorylation.
a.
Serine and threonine protein kinases add phosphate onto a serine or
threonine residues in proteins
b. Tyrosine kinases add phosphate onto tyrosine residues in proteins.
~2000 different protein kinases
Protein phosphatases
Remove phosphate groups from proteins
~300 different protein phosphatases.
90
BIOL 130
MODULE 5
91
Two main types of G proteins
Heteromeric G protein
Consists of 3 different polypeptide subunits: , , (Fig. 15.4)
Cycles between two states
Active state: GTP is bound to subunit.
Inactive state: GDP is bound to subunit.
These two states cycle back and forth.
Monomeric G protein (e.g. Ras)
Cycles between two states
Active: GTP bound
Inactive: GDP bound
Accessory proteins that regulate G proteins (Karp 628; Fig. 15.19)
a.
GTPase-activating proteins (GAPs)
Stimulate hydrolysis of bound GTP.
This inactivate G protein.
b. Gaunine nucleotide-exchange factors (GEFs)
Stimulate release of GDP from G protein.
This allows binding of GTP and G protein activation.
c.
Gaunine nucleotide-dissociation inhibitors (GDIs)
GDIs inhibit release of GDP from G protein.
This maintains G protein in inactive state (GDP bound).
5a 3) Two alternative types of signal transduction systems (Karp p. 607; Fig. 15.2)
1.
The receptor is coupled to heterotrimeric G protein (see section below).
> 100 different G protein-coupled receptors have been identified.
G protein coupled receptors (GPCRs) contain 7 transmembrane helices.
- has diffusible second messenger.
2.
Receptor is not coupled to G-proteins but instead has tyrosine kinase activity.
Receptor Tyrosine Kinases (RTKs)
> 50 different RTKs have been identified (deal with in Unit 23)
5a 4) Interaction of heteromeric G protein with receptor and effector (Karp: p. 609, Fig. 15.4)
1.
2.
3.
4.
Upon binding ligand, receptor then binds G protein.
This causes subunit to exchange GDP for GTP.
In turn, this causes subunit to separate from other subunits.
subunit then binds to the effector, activating the effector (see section below).
BIOL 130
MODULE 5
92
Two effectors activated by G protein
1.
2.
Adenylyl (or adenyl) cyclase (Karp: p. 630, Fig. 15.11)
Converts ATP into a second messenger product: cAMP (see section I below).
Phospholipase C (Karp p. 626, Fig. 15.7) (deal with in Unit 22)
Splits phosphatidylinositol 4,5 bisphosphate (PIP2) into two second
messenger products:
Diacylglycerol (DAG)
Inositol 1,4,5-trisphosphate (IP3)
5a 5) cAMP
Most famous 2nd messenger
Exerts its effect by activating protein kinase A (PKA).
PKA
A cAMP-dependent protein kinase.
cAMP binds to allosteric site on regulatory subunit.
This releases inhibition by regulatory subunit.
PKA phosphorylates serine and threonine on a wide variety of different proteins
(Karp: p. 621 Fig. 15.13)
Mediates many hormone-induced responses. (Karp p. 620 Table 15.4)
Example of response induced by cAMP: glucose mobilization (see section M below).
Glucose mobilization and cAMP (Figs. 15.10, 15.11, 15.12)
1.
2.
3.
4.
5.
6.
7.
8.
9.
Epinephrine binds receptor
This activates heteromeric G protein.
This activates adenyl cyclase.
This forms cAMP.
cAMP activates PKA.
Two PKA activities regulate glucose mobilization
a.
PKA phosphorylates glycogen synthase, which inhibits it.
(prevents glycogen synthesis from glucose)
b. PKA phosphorylates phosphorylase kinase, which activates it.
Phosphorylase kinase phosphorylates phosphorylase, which activates it.
Phosphorylase breaks down glycogen to glucose
Blood glucose levels rise
BIOL 130
MODULE 5
93
5b: Other Secondary Messengers: Lipids, Calcium and
Nitric Oxide: Unit #21
(Karp: pp. 614-617; 634-638; 640-642)
5b 1) Lipid-derived 2nd messengers (Karp 626; Fig. 15.7)
Phospholipids of cell membranes are a source of 2nd messengers.
Phosphatidylinositol (PI) is best studied source of lipid 2nd messenger.
PI comes in different phosphorylated forms (phosphoinositides).
Converted to 2nd messengers by phospholipases (hydrolytic enzymes).
Phospholipases split phospholipids into their component parts.
an example is phosphatidyl inositol-specific phospholipase C- (PI-PLC)
1. Generation of 2nd messengers from PI (Karp 616; Fig. 15.8)
a.
PI phosphorylated by phosphoinositide kinases.
PI PI(4)P PI(4,5)P2 (or PIP2)
Phosphorylated inositol ring of phosphoinositides form binding sites for certain proteins.
Proteins with PH' domains can bind to these sites.
b.
PI-PLC has a PH domain.
its PH domain holds the enzyme at the inner surface of plasma membrane.
c.
PI-PLC activated by heterotrimeric G protein
Splits PIP2 into
a.
Diacylglycerol (DAG)
b. Inositol 1,4,5-trisphosphate (IP3)
2. diacylglycerol (DAG) ( Fig. 15.8)
Remains in plasma membrane.
Activates protein kinase C.
Protein kinase C phosphorylates serine and threonine residues on a wide variety of
target proteins.
Protein kinase C regulates many processes (Karp p. 617 Table 15.2), including cell
proliferation.
Phorbol esters (plant secondary metabolites) activate protein kinase C and cause
normal cells to act temporarily as cancer cells.
3. Inositol 1,4,5-trisphosphate (IP3) ( Fig. 15.8)
IP3 controls a variety of cellular responses (Karp p. 617 Table 15.3)
After formation at plasma membrane, IP3 diffuses into the cytosol.
Binds to a specific IP3 receptor located at the surface of smooth endoplasmic
reticulum (SER).
This releases calcium from SER into cytosol (see next section).
BIOL 130
MODULE 5
5b 2) Calcium ions as second messengers (Karp: p. 634-637)
Many different stimuli induce their response in a target cell by causing a sudden
increase in the concentration of Ca++ in the cytosol.
Change can be:
i.
Oscillation
e.g. liver cells in response to vasopressin (Fig. 15.9)
ii. Elevation in a local region
e.g. macrophages undergoing phagocytosis
e.g. neurons (Fig. 15.25)
iii. A wave spreading from one end of cell to the other
e.g. egg upon fertilization (Fig. 15.27)
1. [Ca++] in different compartments
1.
2.
3.
Cytosol very low ~10-7 M
Certain organelles very high (10,000 x 10-7 M)
a.
Smooth endoplasmic reticulum (SER)
b. Mitochondria
Extracellular medium very high (10,000 x 10-7 M)
2. Maintaining low cytosol Ca++ levels
1.
2.
3.
4.
Ca++ pump in plasma membrane transports Ca++ out of the cell.
Some cells have sodium-calcium exchangers that further reduce cytosolic Ca++
concentrations.
Ca++ pump in SER sequesters Ca++ in SER lumen.
Mitochondria can transport Ca++ into the mitochondrial matrix.
3. What mechanisms cause cytosolic [Ca++] to increase?
1.
2.
Opening of Ca++ channels in plasma membrane.
e.g. neurons
Release of Ca++ from intracellular stores-usually SER
Two main Ca++ channels present in SER membrane.
a.
IP3 receptor channel in SER
Opened by IP3
b. Ryanodine receptor (RyRs) in SER.
Found primarily in excitable cells.
can be opened by calcium itself.
94
BIOL 130
MODULE 5
4. Regulation of ryanodine receptor (RyRs) (Karp: p. 635 Fig. 15.26)
Opened by:
1. Calcium-induced calcium release (CICR)
e.g. cardiac muscle
a.
action potential opens voltage-gated Ca++ channel
b. causes Ca++ flux
c.
Ca++ flux opens RyRs channel.
d. releases Ca++
d. this triggers cell contraction.
e.
Caffine makes RyRs receptor more sensitive to Ca++ and cells more
excitable.
f.
pumping out Ca++ causes contraction
5. How does the Ca++ act in general?
1. Ca++ binds to a small number of calcium-binding proteins and acts through them.
2. Most widely distributed calcium-binding protein is calmodulin.
3. Calcium-calmodulin complex activates or inhibits various enzyme and transport
systems.
Examples
a.
Protein kinases
b. Activates plasma membrane calcium-transport system (self regulation).
6. Self regulation
Calcium-calmodulin complex activates calcium-transport system of the plasma
membrane to pump Ca++ out and restore low cytosolic levels.
5b 3) Nitric oxide as an intercellular messenger (Karp: pp. 640-642)
1. Nitric oxide or NO
1.
2.
3.
Made by an enzyme called nitric oxide synthase (NOS)
Made in copious amounts by macrophages under some conditions
Used to kill bacteria. (does not normally have signaling role in this situation)
excessive NO can lead to pathological signaling (sepsis)
Made in low levels by endothelial cells (inner surface of blood vessels)
Regulate surrounding smooth muscle cells.
95
BIOL 130
MODULE 5
96
2. NO signal transduction pathway in blood vessels (Fig. 15.34)
1.
2.
3.
4.
5.
6.
7.
Acetylcholine binds outer surface of endothelial cell.
Causes rise in cytosolic Ca++ that activates NOS to make NO.
NO diffuses out of endothelial cells into surrounding smooth muscle cells.
NO binds and stimulates guanylyl cyclases in smooth muscle cells to produce cGMP.
cGMP decreases cytosolic Ca++ levels, which causes relaxation of smooth muscle.
Relaxation causes dilation of blood vessels.
This pathway can be manipulated in medicine (angina, septic shock, penis).
Examples of NO in medicine
1. Angina
Inadequate blood flow to heart.
1. Nitroglycerine is metabolized to NO.
2. NO stimulates relaxation of smooth muscles lining blood vessels of heart.
3. Increases blood flow to heart.
2. Septic shock
Dramatic drop in blood pressure caused by bacterial infection.
1. LPS of bacterial cell wall stimulates excessive NO production.
2. NO leads to widespread dilation of blood vessels.
3. NOS inhibitors might be a treatment.
3. Erections
1.
2.
3.
4.
5.
6.
7.
During sexual arousal, nerve impulses to penis triggers release of NO.
NO stimulates cGMP formation.
cGMP causes relaxation of smooth muscle cells in lining of penile blood vessels
and engorgement of penis with blood.
Viagra inhibits a penile isoform of cGMP phosphodiesterase (PDE5).
Phosphodiesterases break down cGMP.
Inhibiting PDE5 maintains elevated cGMP levels.
This promotes development and maintenance of an erection.
BIOL 130
MODULE 5
5c: Receptor Tyrosine Kinases and Cell Proliferation and
Cell Death: Unit #22
(Karp: pp. 623-630; 638-640; 642-646)
5c 1) Receptor Tyrosine Kinases (RTKs)
RTKs are an example of another major type of signaling pathway.
RTKs are often pathways that lead to cell proliferation.
RTKs have two activities.
a.
As cell surface receptors
b. As protein kinases
RTKs act as receptors on the cell surface for:
a.
b.
Polypeptide growth factors such as
i.
Insulin
ii. Epidermal growth factor (EGF)
iii. Platelet-derived growth factor (PDGF)
Cytokines
Polypeptides secreted by one type of immune cell that elicit a response in
another type of immune cell
Example: interferons
RTKs act as tyrosine kinases.
Tyrosine kinases are enzymes that add phosphates to specific tyrosine residues of
proteins.
Tyrosine kinases are involved primarily in control of cell proliferation and
differentiation (rather than intermediary metabolism)
Over 50 different RTKs have been identified.
RTKs do not phosphorylate every tyrosine in a substrate protein.
RTKs phosphorylate tyrosines within certain amino acid sequences.
These are called phosphotyrosine motifs.
Effectors for RTK signaling pathways
Proteins that contain cell-signaling domains, called SH2 domains.
SH2 domains contain high affinity binding sites for phosphotyrosine motifs.
Interaction between phosphotyrosine motifs and SH2 domain in effector protein
leads to different consequences.
Specific consequence depends on effector.
97
BIOL 130
MODULE 5
98
5c 2) EGF receptor as a RTK (Karp: p. 624 Fig. 15.15)
1.
2.
3.
4.
5.
6.
EGF is a mitogen stimulates cell proliferation.
EGF receptor is a monomer in unstimulated cells.
After EGF binds, the monomers interact to form dimers.
Dimerization activates the tyrosine kinase activity of the EGF receptor.
This leads to phosphorylation of tyrosine residues on the cytoplasmic domains of
the receptor autophosphorylation.
Phosphotyrosine residues serve as binding sites for proteins with SH2 domains.
Activation of Raf (Karp: p. 629 Fig. 15.20)
1.
2.
3.
4.
Grb2:
i.
A specific SH2 protein. (Karp: p. 626 Fig. 15.17a; p. 629 Fig. 15.20)
ii. Functions as an adaptor protein.
iii. One domain binds to phosphorylated EGF receptor.
iv. Another domain binds to and activates Sos.
Sos:
i.
Is a GEF (help activate G proteins).
ii. Activates specifically a G protein called Ras.
Ras:
i.
A monomeric G protein
ii. Ras-GTP recruits another protein called Raf to plasma membrane.
Raf:
i.
A protein kinase
ii. At plasma membrane becomes activated protein kinase.
iii. Raf initiates MAP kinase cascade.
The MAP kinase cascade (mitogen-activated protein kinase) (Karp: p. 629 Fig. 15.20)
1.
2.
3.
4.
An orderly chain of phosphorylation reactions initiated by Raf (MAP kinase kinase
kinase).
The last protein kinase in the cascade (MAPK) enters the nucleus and
phosphorylates specific transcription factors.
This increases the affinity of transcription factors for regulatory sites on DNA
This allows expression of genes that encode:
a.
Proteins that lead to initiation of DNA synthesis
e.g. c-fos & c-jun.
b. MAPK phosphatase (MPK-1)
Inactivates MAPK
Stops further signaling along pathway.
BIOL 130
MODULE 5
5c 3) Crosstalk between signaling pathways (Karp: p. 638-640 Fig. 15.33)
Signaling pathways resemble an interconnected web.
e.g. cAMP
Mobilizes glucose.
Also inhibits proliferation of many cells.
cAMP inhibits growth by
i.
Activating PKA
ii. PKA phosphorylates Raf
iii. Phosphorylation inhibits Raf and thus MAP kinase cascade.
5c 4) Pathways that lead to cell death (apoptosis) (Karp: p. 642-646)
Apoptosis
1.
2.
3.
A type of orderly or programmed cell death. (Fig. 35)
Cell responds to certain signals by committing suicide.
Performs tissue remodeling in embryo
e.g. cells are killed selectively in paddle like region that will become hand
Has several important functions in adults
a.
Eliminate specific cells
e.g. activated T lymphocytes that are no longer required.
b. Eliminate cells that have sustained irreparable genetic damage.
c.
Eliminate excess cells to maintain tissue size
(i.e. balance to cell proliferation)
Loss of capacity to undergo apoptosis is a step to becoming cancerous.
Caspases
Special proteolytic enzymes involved in apoptosis.
Exist in an inactive or weakly active form procaspases.
Activated to caspases.
Different caspases are involved in different phases of apoptosis.
Activation phase of apoptosis.
Cell responds to "death signals" (external or internal stimuli)
Involves "initiator" caspases (caspases 8 & 9).
Execution phase of apoptosis.
Death sentence is carried out.
Uses "executioner" caspases.
99
BIOL 130
MODULE 5
100
5c 5) External stimuli for activation phase of apoptosis
1.
2.
Removal of growth factors
e.g. testosterone and prostate cells
Certain proteins
e.g. tumor necrosis factor (TNF) produced by immune cells in response to viral
infection
Receptor-mediated (extrinsic) pathway for apoptosis by external stimuli (Fig. 15.36)
e.g. TNF
1. TNF binds to a transmembrane receptor (TNFR1)
2. Cytoplasmic side contains death domains.
3. Binding of TNF to TNFR1 changes conformation of death domains.
4. Death domains recruit a number of proteins to form complex.
5. Last proteins to join are two procaspase-8 molecules.
6. They activate each other to caspase-8.
7. Caspase-8 (initiator caspase) activates executioner caspases.
8. Executioner caspases bring about apoptosis.
5c 6) Internal stimuli for activation phase of apoptosis
1.
2.
3.
Irreparable genetic damage
High concentrations of cytosolic Ca++.
Severe oxidative stress (ROS = reactive oxygen species)
i.e. free radicals
Mechanism for apoptosis by internal stimuli (intrinsic or mitochondrial pathway)
(Fig. 15.37)
1.
2.
3.
4.
5.
Activate proapoptotic cytoplasmic factors
e.g. some members of Bcl-2 family (Bad, Bid & Bax)
Bad attaches to outer mitochondrial membrane.
Promotes release of cytochrome C from intermembrane space into cytosol.
Cytochrome C forms multiprotein complex with and activates procaspase-9.
Caspase-9 (initiator caspase) activates executioner caspases.
Executioner caspases bring about apoptosis.
BIOL 130
MODULE 5
5c 7) Execution phase of apoptosis
1.
Targets for destruction by executioner caspases
a.
Protein kinases
e.g. Focal adhesion kinase (FAK)
Leads to detachment of apoptotic cell from neighbors.
b. Lamins
Leads to disassembly of nuclear lamina and shrinkage of nucleus.
c.
DNA repair enzymes
2.
Targets for activation by caspases
CAD caspase activated DNase
An endonuclease
Cleaves DNA to 180 base pair fragments
3.
"eat me" signals appear at cell surface.
e.g. phosphatidyl serine
Phagocytes eat or engulf apoptotic cells. (Fig 15.38)
5c 8) Antiapoptotic mechanisms
1.
2.
3.
Opposing signals that maintain cell survival.
Mediated by antiapoptotic members of Bcl-2 family
e.g. Bcl-2 and Bcl-X,
Bind to outer mitochondrial membrane and block release of cytochrome C.
Therefore, abnormal Bcl-2 and Bcl-X are oncogenes. (cause cancer).
101
BIOL 130
MODULE 6
Module 6 outline and notes: Biology of Cancer
6a Regulation of cell proliferation gone wrong
6a1) Introduction to Cancer
6a2) Phenotype of a Cancer cell
6a3) Causes of Cancer
6a4) Genetics of Cancer
6a5) Two types of genes implicated in causing cancer
6a6) Examples of tumor suppressor genes
6a7) Proto-oncogenes vs oncogenes
6a8) Examples of oncogenes
6a9) Overview of several signal pathways involved in tumorigenesis
6b Regulation of cellular social behavior gone wrong
6b1) Metastasis
6b2) Glycocalyx or cell coat
6b3) Extracellular matrix (ECM)
6b4) Macromolecules of ECMs
6b5) Degradation of ECM
6b6) Integrins: the most important family of receptors that attach cells to ECM
6b7) Adhesion of cells to noncellular substrates
6b8) Adhesion of cells to other cells
6b9) Metastasis and anticancer therapy
6b10) Future
102
BIOL 130
MODULE 6
103
6a: Regulation of cell proliferation gone wrong: Unit #23
(Karp: pp. 650-671)
6a 1) Introduction to Cancer
is a disease involving abnormalities in cellular control mechanisms.
Cells proliferate when they shouldn't.
Cells move to where they don't belong. (This is metastasis, next unit)
Tumor mass of cells
i.
Benign tumor cease to grow after reaching a certain size.
ii. Malignant tumor cells divide indefinitely and metastasize.
6a 2) Phenotype of a Cancer Cell
A.
Growth properties
1.
3.
4.
Contact-inhibition of growth (Fig. 16.3)
Property of normal cells but not tumor cells.
Anchorage dependent growth
Property of normal cells but not tumor cells
Tumor cells are usually immortal, but not normal cells.
Cancer cells require fewer growth factors. (Fig. 16.4)
B.
Other differences between normal vs. tumor cells
1.
Cytoskeleton is less well organized in tumor cells. 2. Contact-inhibition of motility
Normal cells stop moving when bump into neighbors but not tumor cells
Cancer cells often have abnormal chromosome numbers.
Aneuploid (Fig. 16.5)
Tumor associated antigens
New cell surface proteins are expressed on some cancer cells.
2.
3.
4.
6a 3) Causes of cancer
1. Chemical carcinogens
cause mutations in DNA of somatic cells.
2. Irradiation
cause mutations in DNA of somatic cells.
3. Oncogenic viruses
Increases risk rather than being sole determinant.
Associated with only a small fraction of human cancers.
6a 4)Genetics of cancer
1. Cancer is monoclonal.
BIOL 130
2.
3.
MODULE 6
104
Caused by uncontrolled proliferation of a single wayward cell.
Tumorigenesis
A multistep process characterized by a progression of genetic alterations in a
single line of cells.
Genetic alterations makes cells increasingly
a.
Less responsive to body's normal regulatory machinery
b. and better able to invade normal tissues (Fig. 16.8).
6a 5) Two types of genes have been implicated in causing cancer (carcinogenesis).
1. Tumor-suppressor genes (Karp: p. 659 Fig. 16.10)
Encode proteins that restrain cell growth and prevent cells from becoming
malignant.
Examples are Rb and p53.
Suffer loss-of-function mutations renders them unable to restrain cell growth.
2. Oncogenes
Encode proteins that promote the loss of growth control and acquisition of
malignancy. (Karp: p. 686)
Gain-of-function mutations that lead to cancer.
6a 6) Examples of tumor suppressor genes
1. RB codes for retinoblastoma (Rb) protein (Fig. 16.11)
a. Rb protein plays a key role in governing cell cycle progression (G1/S).
b. Rb does this by regulating transcription factor E2F
c. E2F regulates synthesis of proteins required for S phase.
d. dissociation of Rb from E2F is controlled by Rb state of phosphorylation.
e. hypophosphorylated Rb binds E2F and restricts proliferation.
f. hyperphosphorylated Rb dissociates from E2F and permits proliferation.
2. p53 (Fig. 16. 13)
a. p53 = polypeptide with a molecular weight of 53 kilodaltons.
b. p53 is sometimes called guardian of genome.
c. Normally after DNA damage, p53 can cause
i. Cell cycle arrest allowing DNA repair
ii. Or apoptosis.
d. If both copies of p53 gene are inactivated, cell either:
i. Dies from mitotic failure or
ii. Proliferates with genetic abnormalities leading to tumor cell formation.
Regulation of p53 activity (Fig. 16.16)
a.
level of p53 in a healthy G1 cell is kept very low by another protein MDM2.
b. level of p53 rises rapidly in G1 cells with DNA damage.
c.
ATM detects DNA damage and phosphorylates p53.
d. Phosphorylated p53 no longer binds inhibitor protein MDM2.
BIOL 130
e.
f.
MODULE 6
105
p53 now accumulates and is an active transcription factor.
p53 up regulates
i. p21, which causes cell cycle arrest
ii. or bax, which causes apoptosis.
3. BRCA1 and BRCA2
- code for proteins that are part of a multiprotein complex involved in DNA repair.
- responsible for most inherited cases of breast cancer
4. PTEN
- lipid phosphatase.
- converts PIP3 to PI (4,5) P2 by removing a phosphate.
- mutation in PTEN leads to excessive PIP3.
- PIP3 activates PKB (Akt), which is a serine-threonine kinase.
- activation of PI3KPKB pathway leads to an increased likelihood that a cell will survive
a stimulus that normally would lead to its destruction.
6a 7) Proto-oncogenes vs. oncogenes
Proto-oncogenes
1. Normal genes that encode proteins having roles in the normal activities of cells.
2. Oncogenes arise from proto-oncogenes through mutation (Karp: p. 679 Fig. 16.12)
3. Oncogenes have been acquired by some viruses, oncogenic viruses.
4. Most oncogenes are derived from proto-oncogenes that play a role in the pathways
that transmit growth signals from the extracellular environment to the cell interior,
particularly the cell nucleus. (Karp: p. 679 Fig. 16.20)
6a 8) Examples of oncogenes
1. Oncogenes that encode growth factors (Karp: p. 679)
Example: sis carried by the cancer-causing simian sarcoma virus.
Codes for platelet-derived growth factors.
Excess PDGF causes cells to proliferate in an uncontrolled fashion.
implicated in some brain tumors.
2. Oncogenes that encode growth factor receptors (Karp: p. 664)
Example: erbB carried by avian erythroblastosis virus.
Codes for altered EGF receptor that continuously expresses tyrosine kinase
activity.
Cells grow independent of the growth factor.
implicated in lung cancers from patients who never smoked.
3. Oncogenes that encode cytoplasmic protein kinases
Example: raf mutation in raf gene that causes Raf (a serine-threonine kinase) to be
permanently active as a kinase.
This leads to continual activation of the MAP Kinase cascade and of cell
proliferation.
4. Oncogenes that encode nuclear transcription factors
BIOL 130
MODULE 6
106
Example: MYC best studied oncogene whose product acts as a transcription
factor.
Myc regulates expression of genes necessary for Go to G1 transition.
Over expression of Myc leads to cell proliferation.
MYC is one of the proto-oncogenes most commonly altered in human cancers.
5. Oncogenes that encode products that affect apoptosis
Any alteration that reduces a cell's ability to self-destruct increases the likelihood of
that cell giving rise to a tumour.
Example: BCL-2 encodes a membrane-bound protein that inhibits apoptosis.
BCL-2 becomes oncogenic when bcl-2 is made at higher than normal levels
(over expressed).
BCL-2 over expression is thought to be cause of certain lymphoid cancers.
6a9) Overview of several processes and signal pathways involved in tumorigenesis
(Karp p 666, Fig 16.18)
1. apoptosis
key inhibitors:
a. Bcl-2
b. PKB
key activators
a. p53
2. cell cycle progression
key activators
a. Raf
b. E2F
3. immortalization
key activator
a. Myc
BIOL 130
MODULE 6
107
6b: Regulation of cellular social behavior gone wrong: Unit #24
(Karp: pp. 230-258; 675-676)
6b 1) Metastasis
- the spread of a tumor within the body.
- if cancer cells only proliferated abnormally, they could be surgically removed.
- instead they leave the primary tumor mass and initiate secondary tumors.
- therefore, the normal interactions between cells and between cells and the extracellular
matrix need to be known.
6b 2) Glycocalyx or cell coat
1.
2.
3.
4.
5.
Carbohydrate layer on surface of animal cells.
Extends outward from plasma membrane.
Made up of carbohydrate units of membrane glycolipids and glycoproteins.
Literally "sugar coating".
General glyocalyx functions:
a.
Reception of outside stimuli
b. Receptors in signal transduction systems (see Unit #21)
6b 3) Extracellular matrix (ECM)
1.
2.
3.
Many cells secrete and organize materials into region beyond plasma membrane.
In multicellular animals, this is ECM.
ECM has two general functions.
a.
Support and protective material.
b. Permit expression of differentiated cell functions.
ECMs of some animal tissues
A.
Epithelial tissue
1. Basal surface of epithelial tissues sits on basement membrane or basal lamina.
2. Basal lamina:
a.
Is 50 to 200 nm thick.
b. Acts as a barrier to passage of macromolecules.
c.
Is doubled layered in kidney.
B.
Connective tissue
e.g. cartilage, bone, tendons
ECM dominates and gives tissues identifiable properties.
e.g. collagen in tendons and proteoglycans in cartilage
6b 4) Macromolecules of ECMs (Karp: p. 233 Fig. 7.5)
1. Collagens
BIOL 130
MODULE 6
108
Glycoproteins abundant in ECM.
All collagens consist of 3 polypeptide chains.
Chains called chains but they can be different.
Greater than 15 distinct types of collagen.
Some collagens (I, II, & III) are fibrillar.
They assemble into cablelike fibrils.
Collagen IV is nonfibrillar.
Each collagen restricted to particular locations within body. e.g. collagen IV in all
basement membranes.
2. Proteoglycans (Karp: p. 236 Fig. 7.9)
Protein-polysaccharide complex.
Consists of core protein to which chains of glycosaminoglycans (GAGs) are attached.
GAGs made up of repeating disaccharide in which two sugars are different.
GAGs are highly acidic due to sulfate and carboxyl groups attached to sugar rings.
Negative charges attract cations, which attract water.
Proteoglycans form a porous, hydrated gel that acts like 'packing material'.
3. Noncollagenous glycoproteins of ECM
These are important in:
a.
Embryonic development
b. Metastasis spread of tumor cells.
a.
Fibronectin (Karp: p. 237 Fig. 7.10)
Two important sites or domains
a.
Collagen-binding domain
b. Cell-binding domain (RGD-containing loop)
b.
Laminin (Karp: p. 239 Fig. 7.12)
Binds to cell surface receptors.
Interwoven with collagen IV into a porous scaffolding.
6b 5) Degradation of ECM
1.controlled ECM degradation is required for normal development & tissue
remodeling . e.g. formation of blood vessels -angiogenesis.
2.also involved in metastasis.
3.-accomplished largely by a family of zinc-containing enzymes, matrix
metalloproteinases (MMPs).
4.MMPs are either secreted or anchored to the plasma membrane.
5.cancer cells induce the synthesis and secretion of MMPs by the surrounding cells.
6.protein fragments of ECM act back on cancer cells to stimulate or inhibit their
growth & invasion.
6b 6) Integrins: the most important family of receptors that attach cells to ECM
(Karp: p. 240-241; Figs 7.13; 7.14)
A superfamily of integral membrane proteins.
BIOL 130
MODULE 6
109
Composed of 2 membrane-spanning polypeptide chains, and .
15 different chains and 8 different chains.
Of > than 100 possible pairings ~20 different integrins have been identified.
Cytoplasmic domain binds cytoskeleton.
Extracellular portion contains binding sites for ECM ligands.
~ of all integrins contain an RGD-binding site.
Binding of all ligands to integrins requires Ca++ or Mg++.
Integrins contain distinct Ca++ binding sites.
Integrin activities (Figs 7.14; 7.15; 7.17c)
A. Adhesion functions
1. adhesion of cells to ECM or substratum
2. adhesion of cells to other cells less common
B. Tansmission of signals
1. Outside-in signaling
- binding to extracellular ligand can cause conformational change in the cytoplasmic
end of the integrin.
This can activate FAK (focal adhesion kinase) and start signal cascade (Fig.
7.17c).
2. inside-in signaling
- many integrins exist on the cell surface in an inactive conformation.
- events within cytoplasm can alter the conformation of the cytoplasmic domain
of the integrin subunits.
- this increase the binding of integrin to an extracellular ligand (Fig. 7.14).
e.g. when blood clotting is needed, IIb integrins on platelets are activated to
bind fibrinogen and form a clot (Fig 7.15)
6b 7) Adhesion of cells to noncellular substrates ( Fig. 7.17)
Two specialized adhesive structures anchor cells to their substratum.
Both involve integrins.
Focal adhesions
Hemidesmosomes
Focal adhesions or focal contacts are found in cells in culture dishes. (Fig. 7.17)
Scattered sites where cells come into close apposition (~10nm) to surface of dish.
Integrins are clustered at focal adhesions.
These connect to material that coats culture dish.
Cytoplasmic domain of these integrins link to microfilaments (actin).
Hemidesmosomes
BIOL 130
MODULE 6
110
Are found in body between epithelial cells and underlying basement membrane.
Contain a dense plaque on inner surface of plasma membrane. (Karp: p. 244 Fig. 7.19)
Intermediate filaments (keratin) course out from the plaque.
Keratin filaments are linked to ECM by integrins.
6b 8) Adhesion of cells to other cells
Four distinct families of membrane proteins mediate cell-cell adhesion. (Karp: p. 253 Fig.
7.28)
Also have potential to carry out transmembrane signaling.
Selectins
Certain members of immunoglobulin superfamily (IgSF)
Certain members of integrin superfamily
Cadherins
Selectins (Karp: p. 2246 Fig. 7.21)
Cell-adhesion molecules that recognize a grouping of 4 carbohydrates.
The carbohydrates are cell surface glycolipids and glycoproteins.
Three known selectins
E-selectin is on endothelial cells.
P-selectin is on platelets and endothelial cells .
L-selectin is on all types of white blood cells (leukocytes).
Selectins mediate transient interactions between circulating leukocytes and vessel
walls. (Karp: p. 259 Fig. 1).
Immunoglobulin superfamily (IgSF) (Karp: p. 249 Fig. 7.22)
Includes soluble immunoglobulins.
But most members are integral membrane proteins on surface of lymphocytes.
Some of these mediate Ca++-independent cell-cell adhesion.
Mediate specific interactions of lymphocytes with cells required for an immune
response. (e.g. macrophages or other lymphocytes).
Some IgSF members are on other cell types
e.g. NCAM & L1 on neurons
e.g. ICAM & VCAM on endothelial cells
Integrins in cell-cell adhesion
A few integrins mediate cell-cell adhesion by binding to IgSF proteins on opposing
cells. (Karp: p. 253 Fig. 7.28).
Example is binding of leukocytes to endothelial cells.
endothelial cells express ICAM.
Neutrophils have an integrin that recognizes ICAM.
BIOL 130
MODULE 6
111
e.g. integrin-IgSF interaction.
helps move neutrophils from bloodstream into tissues during inflammation. (Karp:
p. 247 Fig. 1).
Cadherins (Karp: p. 250 Fig. 7.23)
Family of ~ 12 related glycoproteins that mediate Ca++-dependent cell adhesion.
Cadherins join cells of similar type to one another.
Do so by binding to same cadherin present on surface of neighboring cell.
Some examples are E-cadherin (epithelial) and N-cadherin (neural).
Typically distributed diffusely all along the surfaces of 2 adjacent cells.
Also participate in formation of specialized intercellular junctions
(adherens junctions & desmosomes- see section 6b9)
Cadherins and cancer
-Loss of cadherin function may play a role in the spread of malignant tumors.
-epithelial cell tumors (breast, prostate & colon) have reduced levels of E-cadherin.
- the greater the cadherin reduction, the greater the cells metastatic potential.
6b 9) Metastasis and anticancer therapy
(Karp, p 675-676)
one example
Angiogenesis = formation of blood vessels.
Inhibiting angiogenesis
Growing tumor stimulates formation of new blood vessels (Fig. 16.22).
Blood vessels supply tumor with nutrients and a route to spread.
Tumor cells secrete growth factors for endothelial cells.
Endothelial cells form inner surface of blood vessels.
There are synthetic and natural inhibitors of endothelial cell growth.
e.g. thalidomide and endostatin
Endostatin is proteolytic cleavage product of collagen.
Treating with endostatin caused tumor regression.
6b 10) Future
Trying to understand and treat cancer has led to tremendous gains in basic
knowledge about cells.
These have been triumphs of human intellectual endeavors.
Still treatments remain elusive for many cancers.
Numerous other diseases also have a cellular basis.
Thus many advances in basic cell biology can be expected in the future.
Some of these hopefully can be exploited for cures.
Your Ph.D. likely will involve cell biology.
BIOL 130
SAMPLE FINAL EXAM
112
Sample Final Exam Questions
The final exam is usually between 100 and 110 multiple choice questions.
1.
Which one of the following structures directly gives rise to a primary lysosome?
A. smooth endoplasmic reticulum
B. peroxisome
C. transition vesicle
D. phagosome
E. Golgi body or complex
2.
Which one of the following enzymes is inhibited by ATP?
If none are inhibited, answer e.
A. phosphofructokinase
B. lactate dehydrogenase
C. phosphenol pyruvate carboxylase
D. DNA polymerase
E. none of the above
3.
Which one of the following sequences would be the correct RNA product from the
transcription of the following DNA sequences? ATCGCCAATATT
A. UAGTGGUUAUGG
B. ATCGCCAATATT
C. UAGCGGUUAUAA
D. URGCGGUURURR
E. none of the above
4.
Which one of the following completions is correct?
In a eukaryotic cell RNA polymerase II
A. replicates ribosomal genes
B. transcribes tRNA genes
C. transcribes ribosomal genes
D. translates ribosomal genes
E. transcribes genes for enzymes and structural proteins
5.
Receptor Tyrosine Kinases act as receptors for which one of the following ligands?
A. Low Density Lipoprotein (LDL)
B. insulin
C. epinephrine
D. adenylyl cyclase
E. none of the above
BIOL 130
6.
SAMPLE FINAL EXAM
113
Which one of the following pairs contains both a primary messenger and a
secondary messenger?
A.
B.
C.
D.
E.
cAMP and IP3
DAG and calcium
EGF (epidermal growth factor) and LDL (Low density lipoprotein)
DAG and insulin
EGF and insulin
7.
Which one of the following proteins is responsible for release of coated vesicles
from the membrane?
A. adaptin
B. histone
C. clathrin
D. dynamin
E. collagen
8.
Which one of the following words or phrases best completes the following statement?
In the Z pathway, H+ move from the ______________ to the ______________.
A. inner compartment of thylakoids; stroma.
B. outer chloroplast membrane; cytoplasm.
C. matrix; intermembrane space.
D. stroma; inner compartment of thylakoids.
E. matrix; stroma.
9.
If upon being placed in a solution a plant cell became turgid, what term would
describe the solution?
A. isotonic
B. isoosmotic
C. hypotonic
D. hypertonic
E. If none of the above are correct, answer e.
10. Which of the following completions best fills in the three blanks in the statement below?
A DNA sequence that directs the synthesis of _________________ or of a functional RNA
sequence is defined as a _________________ and would be known to ________________.
A. an enzyme; exon; Beadle and Tatum
B. a transcription factor; promoter; modern cell biologists
C. a polypeptide; gene; modern cell biologists
D. a polypeptide; intron; modern cell biologists
E. an enzyme; cistron; Mr. Bean
BIOL 130
SAMPLE FINAL EXAM
114
Answers To Sample Final Exam Questions
1.
2.
3.
4.
5.
6.
7.
8.
9.
10
E
A
C
E
B
D
D
D
C
C
Try making your own multiple choice questions.
First decide on a fact or concept that you think is worth knowing.
Then decide on how to ask a question that requires knowing this fact or concept in
order to be answered.
One of your choices is the correct answer.
The other choices range from highly unlikely to perhaps one that is close.
Anda mungkin juga menyukai
- Regents Living Environment Practice Questions: New York Regents Living Environment Practice Questions with Detailed ExplanationsDari EverandRegents Living Environment Practice Questions: New York Regents Living Environment Practice Questions with Detailed ExplanationsBelum ada peringkat
- Chem Assignment 2 Christopher BooneDokumen7 halamanChem Assignment 2 Christopher Booneapi-508276223Belum ada peringkat
- Chem 1701 Practice Test 1Dokumen10 halamanChem 1701 Practice Test 1api-535167759Belum ada peringkat
- Chem1701 Assignment1 Winter2020 1Dokumen8 halamanChem1701 Assignment1 Winter2020 1api-403916098Belum ada peringkat
- Channels, Carriers, and Pumps: An Introduction to Membrane TransportDari EverandChannels, Carriers, and Pumps: An Introduction to Membrane TransportBelum ada peringkat
- Biol 1700 Lab 6 Procedure Report FinalDokumen10 halamanBiol 1700 Lab 6 Procedure Report Finalapi-535368507100% (1)
- BIOL 1700 - Test 1 - Important InformationDokumen4 halamanBIOL 1700 - Test 1 - Important InformationSophie100% (1)
- Coursesaver (Chad) College Physics OutlinesDokumen40 halamanCoursesaver (Chad) College Physics Outlinescalong558Belum ada peringkat
- Courseoutline Chem 1701 2018Dokumen19 halamanCourseoutline Chem 1701 2018api-434583946Belum ada peringkat
- Assignment2 Part2Dokumen7 halamanAssignment2 Part2api-438971735Belum ada peringkat
- Sample Exam 4 Fall 11Dokumen14 halamanSample Exam 4 Fall 11janohxBelum ada peringkat
- BIO307 Lecture 5 (Enzyme Kinetics I)Dokumen11 halamanBIO307 Lecture 5 (Enzyme Kinetics I)Phenyo Mmereki100% (1)
- 1 Chemical Kinetics and EquilibriumDokumen77 halaman1 Chemical Kinetics and EquilibriumApril Mergelle Lapuz100% (1)
- BIOC 307 Old Exam 1Dokumen17 halamanBIOC 307 Old Exam 1Katie RoseBelum ada peringkat
- Biochemistry - Final Exam Biology 020.305 December 15, 2011 Instructions For ExaminationDokumen11 halamanBiochemistry - Final Exam Biology 020.305 December 15, 2011 Instructions For ExaminationJenna ReynoldsBelum ada peringkat
- Biochem CombinedDokumen758 halamanBiochem CombinedTheBoss 20Belum ada peringkat
- Lab 4 Chem Brittany GlaspellDokumen11 halamanLab 4 Chem Brittany Glaspellapi-535407136Belum ada peringkat
- Lecture Notes First Semester Yr 2 BPham BMLS BDSDokumen57 halamanLecture Notes First Semester Yr 2 BPham BMLS BDSKarin AdraiBelum ada peringkat
- Biol 1700 Lab 4 Procedure Report FinalDokumen14 halamanBiol 1700 Lab 4 Procedure Report Finalapi-535371469Belum ada peringkat
- Bonding Notes General Chemistry 1Dokumen48 halamanBonding Notes General Chemistry 1JL VABelum ada peringkat
- Mechanism of Enzyme CatalysisDokumen18 halamanMechanism of Enzyme CatalysisBuhlebenkosi Jackie BhebheBelum ada peringkat
- Enzyme: Jahangirnagar UniversityDokumen34 halamanEnzyme: Jahangirnagar UniversityShanian AhmedBelum ada peringkat
- Cell Biology Practice 2Dokumen15 halamanCell Biology Practice 2NgMinhHaiBelum ada peringkat
- scsc7211's Version of Alan's DAT Biology NotesDokumen47 halamanscsc7211's Version of Alan's DAT Biology NotesMusawir YasinBelum ada peringkat
- EnzymesDokumen9 halamanEnzymesramloghun veerBelum ada peringkat
- Enzyme Catalysis LabDokumen8 halamanEnzyme Catalysis LabAzra RamicBelum ada peringkat
- U04 Notes Part4 Intermolecular ForcesDokumen66 halamanU04 Notes Part4 Intermolecular ForcesKhondokar TarakkyBelum ada peringkat
- Exam II Learning Objectives BIOC 384Dokumen30 halamanExam II Learning Objectives BIOC 384amnguye1Belum ada peringkat
- Physical Pharmacy q1 MidtermDokumen78 halamanPhysical Pharmacy q1 MidtermCatherine RiaBelum ada peringkat
- Respiratory Chain & Oxidative PhosphorylationDokumen57 halamanRespiratory Chain & Oxidative PhosphorylationHanifa AffianiBelum ada peringkat
- Cell Organelles ReviewDokumen5 halamanCell Organelles ReviewvictoriaBelum ada peringkat
- Biotechnology 2012 13Dokumen54 halamanBiotechnology 2012 13Aziz KhanBelum ada peringkat
- Unit 5 Chemical KineticsDokumen37 halamanUnit 5 Chemical KineticsSanjay SharmaBelum ada peringkat
- Bio 100 A Virtual Labs Unit OneDokumen7 halamanBio 100 A Virtual Labs Unit OneTammy SmithBelum ada peringkat
- General Physiology: First YearDokumen67 halamanGeneral Physiology: First YearGo HellBelum ada peringkat
- Lecture of Enzymes.Dokumen9 halamanLecture of Enzymes.ibrahim khalilBelum ada peringkat
- Fluid Therapy in Dengue CPG Modify HPP 2017Dokumen37 halamanFluid Therapy in Dengue CPG Modify HPP 2017Nur FadzilahBelum ada peringkat
- Carbohydrates Aka Saccharides NumberDokumen2 halamanCarbohydrates Aka Saccharides NumberAngelica Llamas0% (1)
- EK Schedule - 4 MonthDokumen14 halamanEK Schedule - 4 MonthLauraAlexandraBelum ada peringkat
- Chapter 8 Lecture-Energy, Enzymes, and Metabolism-MODIFIED2Dokumen49 halamanChapter 8 Lecture-Energy, Enzymes, and Metabolism-MODIFIED2E'Lasia LarkinBelum ada peringkat
- A Cell Atlas of Human Thymic Development Defines T Cell Repertoire Formation PDFDokumen12 halamanA Cell Atlas of Human Thymic Development Defines T Cell Repertoire Formation PDFCris BañadosBelum ada peringkat
- AP Psychology Mnomonic DevicesDokumen7 halamanAP Psychology Mnomonic DevicesBellony SandersBelum ada peringkat
- AP Biology Lab ReviewDokumen72 halamanAP Biology Lab ReviewPaweenwat ThongprasopBelum ada peringkat
- Cell Biology LabManual Version 8.0 May 2012 1 TRUNCATEDDokumen57 halamanCell Biology LabManual Version 8.0 May 2012 1 TRUNCATEDதுர்காஸ்ரீ கங்கா ராதிகாBelum ada peringkat
- (1stsem MANUAL) Biochemistry LecDokumen150 halaman(1stsem MANUAL) Biochemistry LecAngela Louise SmithsBelum ada peringkat
- Backwards ReasoningDokumen40 halamanBackwards Reasoningharshit chaudharyBelum ada peringkat
- The Structure and Function of MacromoleculesDokumen70 halamanThe Structure and Function of MacromoleculesLustried Nadyang100% (1)
- Animal Cell Culture BasicsDokumen10 halamanAnimal Cell Culture Basicsசுப விஜய் ப்ரீத்தி100% (1)
- Chapter 2 - Basics of EnzymesDokumen40 halamanChapter 2 - Basics of EnzymesSakinah MuhamadBelum ada peringkat
- Biology Lecture 5 Hormone ChartDokumen2 halamanBiology Lecture 5 Hormone Chartmark_pedersen_6Belum ada peringkat
- Lecture Notes Lectures 1 9 CompleteDokumen244 halamanLecture Notes Lectures 1 9 CompleteBirhanu AyenewBelum ada peringkat
- Biochemistry Laboratory Manual: Biological Sciences M114L Spring 2015Dokumen109 halamanBiochemistry Laboratory Manual: Biological Sciences M114L Spring 2015Rx Chau100% (1)
- Stereochemistry QuestionsDokumen7 halamanStereochemistry Questionsalyson_lBelum ada peringkat
- COURSE WORK MOLECULAR BIOLOGY & GeneticsDokumen3 halamanCOURSE WORK MOLECULAR BIOLOGY & Geneticsusaeed00000Belum ada peringkat
- EnzymesDokumen19 halamanEnzymesوليد. وفي الروح0% (1)
- BIO 3200 - Human Physiology LCT 1 - Ch1-5Dokumen9 halamanBIO 3200 - Human Physiology LCT 1 - Ch1-5HoreaBelum ada peringkat
- PCOL2012 Unit of Study Outline 2016Dokumen10 halamanPCOL2012 Unit of Study Outline 2016ncisisthebestBelum ada peringkat
- BIO307 Lecture 3 (Introduction To Enzymes)Dokumen13 halamanBIO307 Lecture 3 (Introduction To Enzymes)Phenyo MmerekiBelum ada peringkat
- Chapter 7 Cellular RespirationDokumen47 halamanChapter 7 Cellular Respirationwickedbiology101Belum ada peringkat
- Handout Orthopedic Nursing Assisstive DevicesDokumen14 halamanHandout Orthopedic Nursing Assisstive DevicesPaul Christian P. Santos, RN100% (2)
- Cont. Diagnostic Test 2 (61-150)Dokumen13 halamanCont. Diagnostic Test 2 (61-150)Sheila Tolentino-BelanioBelum ada peringkat
- EPA Air Quality Index School Activity ChartDokumen2 halamanEPA Air Quality Index School Activity ChartCourier JournalBelum ada peringkat
- Cleft Lip / Cleft Palate: Predisposing FactorsDokumen15 halamanCleft Lip / Cleft Palate: Predisposing FactorsMabesBelum ada peringkat
- DoH BP Leaflet - Web VersionDokumen2 halamanDoH BP Leaflet - Web Versionpatburchall6278Belum ada peringkat
- 3-Kingdom AnimaliaDokumen16 halaman3-Kingdom AnimaliaEhmz XavièrBelum ada peringkat
- The Transformation of Neuronal Activity Into Conscious Experience: The Syntergic Theory Jacobo GrinbergDokumen10 halamanThe Transformation of Neuronal Activity Into Conscious Experience: The Syntergic Theory Jacobo GrinbergsebastianBelum ada peringkat
- From The Unconscious To The Conscious by Gustave GeleyDokumen348 halamanFrom The Unconscious To The Conscious by Gustave GeleyDonna McNeil100% (3)
- Unit 6 - Topic 6 - Human Physiologyy - (11 HL and SL)Dokumen13 halamanUnit 6 - Topic 6 - Human Physiologyy - (11 HL and SL)John DoeBelum ada peringkat
- PB 840 - Brochure PDFDokumen4 halamanPB 840 - Brochure PDFPriyankaBelum ada peringkat
- Puberphonia 11Dokumen19 halamanPuberphonia 11jaguar1979100% (2)
- Oxidative Phosphorylation V Inhibitors and UncouplersDokumen15 halamanOxidative Phosphorylation V Inhibitors and UncouplersIffatnazBelum ada peringkat
- STM123 - General BiologyDokumen126 halamanSTM123 - General BiologyLEONIEVEVE L LIMBAGABelum ada peringkat
- Difference of Arterial and Venous InsufficiencyDokumen6 halamanDifference of Arterial and Venous InsufficiencyBeep TerradoBelum ada peringkat
- Full Test Kit InfoDokumen232 halamanFull Test Kit InfoolivierBelum ada peringkat
- Cardiovascular 9. Acute Myocardial InfarctionDokumen39 halamanCardiovascular 9. Acute Myocardial Infarctionapi-19641337100% (2)
- RespiratoryDokumen4 halamanRespiratoryHaliana IzatiBelum ada peringkat
- Img023Dokumen1 halamanImg023muthiajayantiBelum ada peringkat
- A Resounding TinkleDokumen73 halamanA Resounding TinkleSula Douglas FolkesBelum ada peringkat
- MegaBrain Volume 2 Number 4Dokumen74 halamanMegaBrain Volume 2 Number 4Frabato Bardon100% (2)
- Biochemistry Quizzes by Ronnie BaticulonDokumen20 halamanBiochemistry Quizzes by Ronnie BaticulonMhartin GarciaBelum ada peringkat
- Singing Technique and Anatomy Part 1Dokumen5 halamanSinging Technique and Anatomy Part 1musicguy joeBelum ada peringkat
- Life ProcessesDokumen36 halamanLife Processes2erwr100% (1)
- PHS 205 Cardiovascular SystemDokumen33 halamanPHS 205 Cardiovascular Systemdivineraymond34Belum ada peringkat
- Musculoskeletal XXXXXXXXXDokumen25 halamanMusculoskeletal XXXXXXXXXIgnatius Erik Dwi WahyudiBelum ada peringkat
- Effect of Omeprazole On Oral and Intravenous RS-Methadone Pharmacokinetics and Pharmacodynamics in The RatDokumen12 halamanEffect of Omeprazole On Oral and Intravenous RS-Methadone Pharmacokinetics and Pharmacodynamics in The Ratdian oktavianiBelum ada peringkat
- Learning Objectives Identify The 6 Classes ofDokumen21 halamanLearning Objectives Identify The 6 Classes ofhahmed78Belum ada peringkat
- Role of Trace Minerals in Reproduction of Dairy AnimalDokumen27 halamanRole of Trace Minerals in Reproduction of Dairy AnimalDr. Umesh Sontakke83% (6)
- A Systematic Approach To The Unconscious PatientDokumen5 halamanA Systematic Approach To The Unconscious PatientDoctor RadiologistBelum ada peringkat
- Love Life: How to Raise Your Standards, Find Your Person, and Live Happily (No Matter What)Dari EverandLove Life: How to Raise Your Standards, Find Your Person, and Live Happily (No Matter What)Penilaian: 3 dari 5 bintang3/5 (1)
- Summary: The Psychology of Money: Timeless Lessons on Wealth, Greed, and Happiness by Morgan Housel: Key Takeaways, Summary & Analysis IncludedDari EverandSummary: The Psychology of Money: Timeless Lessons on Wealth, Greed, and Happiness by Morgan Housel: Key Takeaways, Summary & Analysis IncludedPenilaian: 4.5 dari 5 bintang4.5/5 (82)
- The Age of Magical Overthinking: Notes on Modern IrrationalityDari EverandThe Age of Magical Overthinking: Notes on Modern IrrationalityPenilaian: 4 dari 5 bintang4/5 (32)
- LIT: Life Ignition Tools: Use Nature's Playbook to Energize Your Brain, Spark Ideas, and Ignite ActionDari EverandLIT: Life Ignition Tools: Use Nature's Playbook to Energize Your Brain, Spark Ideas, and Ignite ActionPenilaian: 4 dari 5 bintang4/5 (404)
- ADHD is Awesome: A Guide to (Mostly) Thriving with ADHDDari EverandADHD is Awesome: A Guide to (Mostly) Thriving with ADHDPenilaian: 5 dari 5 bintang5/5 (3)
- By the Time You Read This: The Space between Cheslie's Smile and Mental Illness—Her Story in Her Own WordsDari EverandBy the Time You Read This: The Space between Cheslie's Smile and Mental Illness—Her Story in Her Own WordsBelum ada peringkat
- Raising Mentally Strong Kids: How to Combine the Power of Neuroscience with Love and Logic to Grow Confident, Kind, Responsible, and Resilient Children and Young AdultsDari EverandRaising Mentally Strong Kids: How to Combine the Power of Neuroscience with Love and Logic to Grow Confident, Kind, Responsible, and Resilient Children and Young AdultsPenilaian: 5 dari 5 bintang5/5 (1)
- The Comfort of Crows: A Backyard YearDari EverandThe Comfort of Crows: A Backyard YearPenilaian: 4.5 dari 5 bintang4.5/5 (23)
- The Ritual Effect: From Habit to Ritual, Harness the Surprising Power of Everyday ActionsDari EverandThe Ritual Effect: From Habit to Ritual, Harness the Surprising Power of Everyday ActionsPenilaian: 4 dari 5 bintang4/5 (4)
- When the Body Says No by Gabor Maté: Key Takeaways, Summary & AnalysisDari EverandWhen the Body Says No by Gabor Maté: Key Takeaways, Summary & AnalysisPenilaian: 3.5 dari 5 bintang3.5/5 (2)
- Think This, Not That: 12 Mindshifts to Breakthrough Limiting Beliefs and Become Who You Were Born to BeDari EverandThink This, Not That: 12 Mindshifts to Breakthrough Limiting Beliefs and Become Who You Were Born to BePenilaian: 2 dari 5 bintang2/5 (1)
- Summary: Outlive: The Science and Art of Longevity by Peter Attia MD, With Bill Gifford: Key Takeaways, Summary & AnalysisDari EverandSummary: Outlive: The Science and Art of Longevity by Peter Attia MD, With Bill Gifford: Key Takeaways, Summary & AnalysisPenilaian: 4.5 dari 5 bintang4.5/5 (42)
- Dark Psychology & Manipulation: Discover How To Analyze People and Master Human Behaviour Using Emotional Influence Techniques, Body Language Secrets, Covert NLP, Speed Reading, and Hypnosis.Dari EverandDark Psychology & Manipulation: Discover How To Analyze People and Master Human Behaviour Using Emotional Influence Techniques, Body Language Secrets, Covert NLP, Speed Reading, and Hypnosis.Penilaian: 4.5 dari 5 bintang4.5/5 (110)
- The Courage Habit: How to Accept Your Fears, Release the Past, and Live Your Courageous LifeDari EverandThe Courage Habit: How to Accept Your Fears, Release the Past, and Live Your Courageous LifePenilaian: 4.5 dari 5 bintang4.5/5 (254)
- Raising Good Humans: A Mindful Guide to Breaking the Cycle of Reactive Parenting and Raising Kind, Confident KidsDari EverandRaising Good Humans: A Mindful Guide to Breaking the Cycle of Reactive Parenting and Raising Kind, Confident KidsPenilaian: 4.5 dari 5 bintang4.5/5 (170)
- Manipulation: The Ultimate Guide To Influence People with Persuasion, Mind Control and NLP With Highly Effective Manipulation TechniquesDari EverandManipulation: The Ultimate Guide To Influence People with Persuasion, Mind Control and NLP With Highly Effective Manipulation TechniquesPenilaian: 4.5 dari 5 bintang4.5/5 (1412)
- The Body Keeps the Score by Bessel Van der Kolk, M.D. - Book Summary: Brain, Mind, and Body in the Healing of TraumaDari EverandThe Body Keeps the Score by Bessel Van der Kolk, M.D. - Book Summary: Brain, Mind, and Body in the Healing of TraumaPenilaian: 4.5 dari 5 bintang4.5/5 (266)
- To Explain the World: The Discovery of Modern ScienceDari EverandTo Explain the World: The Discovery of Modern SciencePenilaian: 3.5 dari 5 bintang3.5/5 (51)
- The Obesity Code: Unlocking the Secrets of Weight LossDari EverandThe Obesity Code: Unlocking the Secrets of Weight LossPenilaian: 4 dari 5 bintang4/5 (6)
- Cult, A Love Story: Ten Years Inside a Canadian Cult and the Subsequent Long Road of RecoveryDari EverandCult, A Love Story: Ten Years Inside a Canadian Cult and the Subsequent Long Road of RecoveryPenilaian: 4 dari 5 bintang4/5 (46)
- The Marshmallow Test: Mastering Self-ControlDari EverandThe Marshmallow Test: Mastering Self-ControlPenilaian: 4.5 dari 5 bintang4.5/5 (60)
- Mindset by Carol S. Dweck - Book Summary: The New Psychology of SuccessDari EverandMindset by Carol S. Dweck - Book Summary: The New Psychology of SuccessPenilaian: 4.5 dari 5 bintang4.5/5 (328)
- Why We Die: The New Science of Aging and the Quest for ImmortalityDari EverandWhy We Die: The New Science of Aging and the Quest for ImmortalityPenilaian: 4.5 dari 5 bintang4.5/5 (6)
- How to ADHD: The Ultimate Guide and Strategies for Productivity and Well-BeingDari EverandHow to ADHD: The Ultimate Guide and Strategies for Productivity and Well-BeingPenilaian: 1 dari 5 bintang1/5 (2)