Rains Models PDF
Diunggah oleh
MicheleVonciDeskripsi Asli:
Judul Asli
Hak Cipta
Format Tersedia
Bagikan dokumen Ini
Apakah menurut Anda dokumen ini bermanfaat?
Apakah konten ini tidak pantas?
Laporkan Dokumen IniHak Cipta:
Format Tersedia
Rains Models PDF
Diunggah oleh
MicheleVonciHak Cipta:
Format Tersedia
DRAFT: May 29, 2003
Refinement Application for Inelastic Neutron Scattering
Software and documentation written by R.M. Dimeo
robert.dimeo@nist.gov
Data Visualization and Analysis Team
NIST Center for Neutron Research
CONTENTS
Introduction .............................................................................................................................................................................. 2
Overview of the RAINS environment...................................................................................................................................... 2
Loading DATA in RAINS ....................................................................................................................................................... 3
Data operations/functions......................................................................................................................................................... 4
Model operations/functions...................................................................................................................................................... 6
Data Fitting Example ............................................................................................................................................................... 7
Tunneling functions................................................................................................................................................................ 10
Diffusion functions................................................................................................................................................................. 12
Miscellaneous functions ......................................................................................................................................................... 18
DRAFT: May 29, 2003
Introduction
RAINS is a new application included in the DAVE software package that gives users the power to
analyze their inelastic neutron spectra by fitting models of S(Q,) to their entire data set. In contrast to
another analysis application in DAVE called PAN (Peak Analysis), RAINS is not limited to fitting a
single model to individual detector groups. Rather RAINS fits a parametrized surface given by a
theoretical scattering law S(Q,) to the entire data set. Moreover users can fit different models to the
same data set and compare the least-squares goodness-of-fit metric 2 for instance thus facilitating the
model selection process. Of course this is not a black-box type program so model selection based
solely on 2 can be fraught with peril. Caveat emptor! This document summarizes the features and
functionality of the RAINS program.
Overview of the RAINS environment
RAINS is launched from the main DAVE interface from the menu item labeled Data Analysis a
RAINS. Two detached panels should appear as shown below in Fig. 1. You can move these panels
around independent of one another and resize them at any stage during your data analysis. The panel
on the left contains three regions as shown below. This panel contains a tree hierarchy for ease of file
and model navigation through the fit process and two vertically stacked windows that display the data
and the residual difference between the model and the data. The right panel contains two vertically
stacked windows that display a particular cut or Q value of the data as well as the overall fit. The
residuals are displayed in the bottom window.
Fig. 1 Two-panels in the RAINS fitting environment. See text for an explanation.
The menus in the left panel consist of FILE, MODEL, PRINTING OPTIONS, and HELP selections.
At the moment the PRINTING OPTIONS menu has no items under it as it is a work-in-progress. The
FILE menu contains options to LOAD DAVE (i.e. load a DAVE file) or QUIT. The MODELS menu
DRAFT: May 29, 2003
item contains selections that allow the user to ADD NEW MODEL or select from a further list of
DIFFUSION FUNCTIONS, TUNNELING FUNCTIONS, and/or MISCELLANEOUS FUNCTIONS.
A comprehensive list of these individual functions is shown at the end of this document.
This environment has been designed to enable users to test different models for the same data set in
order to aid in the model selection process. To that end a tree hierarchy has been selected as the main
navigation tool. For each data set users can have as many models as they wish. A single model can be
composed of as many S(Q,) functions as desired. The functions in one model are added together to
form the overall model. The details will be discussed later in this document but a simple example
showing the organization of the functions, models, and data files are shown in Fig. 2. In this figure the
data set labeled 20010621_02dyn.dave has two models associated with it. The first model is composed
of two functions: a background function and a 3-fold jump diffusion model. The second model is
composed of just a 2-fold jump diffusion model. These two models can be fit to the same data set
independently and the goodness-of-fit can be compared.
Fig. 2 Overview of the navigation tool for the function/model/file tree hierarchy.
Loading DATA in RAINS
Any number of existing DAVE files can be loaded into the workspace by selecting the FILE a LOAD
DAVE FILE menu item. Note that the data file must be a function of two variables (Q,E). In general
it is expected that the independent variables are wavevector transfer Q and energy transfer E. In
addition it is expected that the data are uniformly spaced on a regular grid in E. This is checked when
the data file is loaded. Upon successfully loading the files into RAINS new folders will appear on the
tree hierarchy under the DAVE DATA folder and the name of the folders will match the file names.
Moreover a folder labeled MODEL will appear under the new data folder. In addition an image
representation (also called a spectrogram in time-frequency lingo) of the data associated with the
currently selected data folder will appear in the top window of the left panel. An example of a data file
collected on the High Flux Backscattering Spectrometer and reduced using DAVE is shown in Fig. 3.
Note that the cut showing Q=1.51 -1 is displayed in the right panel.
DRAFT: May 29, 2003
Fig. 3 Display resulting from loading a data set collected on HFBS.
You can use the mouse to zoom into the top window of either panel in order to get a magnified view of
your data. This is a standard rubber-band type operation in which you simply drag the cursor over the
plot while holding the left mouse button down. Releasing the left mouse button will execute the
magnification operation. Note that there is an upper bound on the magnification limit.
Data operations/functions
The operations performed within this application are accessible primarily via context-sensitive menus.
These are simply menus that appear when the user presses the right mouse button when the cursor is on
top of one of the folders displayed in the left panel. For instance, when you put the cursor over the
data file folder currently selected and press the right mouse button, you should see the context sensitive
menu like the one shown in Fig. 4.
DRAFT: May 29, 2003
Fig. 4 Context-sensitive menu that appears when user right-clicks the mouse and the cursor is over the data file folder.
Selecting the first choice, ADD A NEW MODEL FOLDER, results in a new folder appearing after the
existing one. You should note that all of the data folders have a single default folder associated with
them. The second choice allows the user to create a Gaussian resolution function based on the data
binning and grouping. Selecting this option launches a utility allowing the user to parameterize the Q
dependence of the full-width-at-half-maximum (FWHM) and the center of the function. This
application is shown in Fig. 5. Note that the user can type in any expression in IDL syntax as a
function of y. Note Y is used in place of Q. The surface is actually represented internally as a function
of the independent variables x and y. Examples of expressions you might put in for the Y-dependence
for the FWHM or center are listed below:
Examples:
Algebraic expression
0.25
0.5+1.05Q2
jo(3.0Q)
Jo(3.0Q)
e-0.5q
IDL syntax
0.25+0.0*y
0.5+1.05*y^2
sph_bessel(3.0*y,0)
beselj(3.0*y,0)
exp(-0.5*y)
Comment
Must include 0.0
When you are satisfied with the resolution function you can press the ACCEPT button and the
resolution function will be present for that data set for the rest of the session. Note that you can create
a new resolution function at any time thereafter by repeating this operation.
DRAFT: May 29, 2003
Fig. 5 Gaussian instrumental resolution function generator application.
Users may also delete any number of data files from memory by selecting the next item in the menu
context-sensitive menu labeled DELETE SELECTED FILE. Finally the user may display descriptive
information related to a particular data file by selecting DISPLAY FILE INFORMATION. The exact
content displayed depends on the instrument from which the data was collected and how the data were
reduced.
Model operations/functions
Another context-sensitive menu appears by left-clicking on the MODEL menu as shown in Fig. 6.
There is a further drop-down menu on this one labeled FITTING (selected in Fig. 6) which contains a
number of selections germane to the fitting process. The first is algorithm selection. For the moment
only the least-squares algorithm is present but work is in progress on incorporating a genetic algorithm
for more difficult fitting problems. The next item in FITTING is SELECT FIT RANGE(S) in which
the user can enter the indices to include in both the Q and E variables for fitting.
DRAFT: May 29, 2003
Fig. 6 Model context-sensitive menu.
Data Fitting Example
In this section we present a demonstration of fitting a tunneling model to real data. The data loaded
into the RAINS environment in Fig. 6 is from the High Flux Backscattering Spectrometer. This
instrument has an approximate Gaussian-like resolution function with a full-width at half maximum of
about 0.8 meV. Therefore we create a resolution function by selecting CREATE GAUSSIAN
RESOLUTION FUNCTION from the data context-sensitive menu as discussed in the section on Data
operations/functions. In the Resolution Function Generator application type 0.8+0.0*y in the field
labeled FWHM (Y) and press carriage return. It is important to press the carriage return so that the
display in the application updates (you should see this update). Next press the ACCEPT button.
Next select the MODEL folder below the data set labeled 20010618_01dyn.dave. Select the MODELS
menu item from the application menu bar, select TUNNELING FUNCTIONS, then 3-FOLD
TUNNELING. A new element labeled 3-FOLD TUNNELING will appear below the MODEL folder
for that data set. Next select the MODEL folder and you should see both the spectrogram and the cuts
windows show the model superposed on the data. RAINS actually tries to guess model parameters
without fitting anything so the resulting plots should look close. An example of what you should see is
shown in Fig. 7.
DRAFT: May 29, 2003
Fig. 7 Initial guess for the model fit.
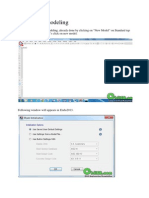
The next step is to set up the fit range. This is accomplished by selecting the MODEL folder and
launching the context-sensitive menu. Select FITTING a SELECT FIT RANGE(S) and you will see
a dialog shown in fig. 8. The existing values for this file are 0-217 for the x-range (energy transfer)
and 0-15 for the y-range (wavevector transfer). Note that these ranges are zero-based indices into the
arrays for each of the independent variables. So in this case there are 218 energy channels and 16
wavevector transfer values. For our purposes we will include all energy transfers in the fit (0-217) but
limit the wavevector transfer to wavevectors with indices between 3 and 14 inclusive. Note that this
dialog supports comma delimited specifications as well. For instance if you wanted to include only
wavevectors 0, 9,10,11,12, and 15 you could type: 0,9-12,15. Press ACCEPT and DISMISS when
you are finished.
Fig. 8 Fit range entry dialog.
DRAFT: May 29, 2003
Now you are ready to fit the model to the data. From the MODEL context-sensitive menu select
FITTING aSTART FITTING. As RAINS fits the data a new dialog display appears that tells the user
how the fit is progressing in terms of the fit iteration, the c2 value, and also provides an interrupt
button which will pause the fit and save the current best-fit parameters at this iteration. This dialog
display is shown in Fig. 9.
Fig. 9 Fit progress dialog.
Once the fit has completed this dialog will disappear. You can examine the resulting fit visually by
looking at individual spectra with the best-fit model superposed as well as view the imagerepresentation of the fit and data. You may examine the parametric results of the fit by selecting
FITTINGaDISPLAY FIT REPORT which launches a dialog that displays the parameters and the
uncertainties based on the fit.
DRAFT: May 29, 2003
Current models included in RAINS
Tunneling functions
3-fold rotational tunneling
Analytic expression:
1 A o (Q)
S(Q, E) = I(Q) A o (Q)(E E c (Q)) +
((E E t E c (Q)) + (E + E t E c (Q)) )
2
where the EISF (elastic-incoherent structure factor) is given by
5 + 4 jo (Qr 3 )
A o (Q) =
9
and jo is the zeroth-order spherical Bessel function.
Model parameters:
I(Q):
Ec(Q):
Et:
r:
group/detector-dependent overall intensity
group/detector-dependent center of scattering function
tunnel splitting
tunnel circle radius ()
DRAFT: May 29, 2003
Broadened 3-fold rotational tunneling
Analytic expression:
S(Q, E) = I(Q)
[A o (Q)(E E c (Q)) +
1 A o (Q)
(L(E E c (Q); q ) + L(E E t E c (Q); t ) + L(E + E t E c (Q); t ) )
3
where the EISF (elastic-incoherent structure factor) is given by
1 + 2 jo (Qr 3 )
A o (Q) =
3
jo is the zeroth-order Bessel function, and the Lorentzian broadened components are given by
1
L(E; ) =
.
2
E + 2
Model parameters:
I(Q):
Ec(Q):
Et:
t:
q :
r:
group/detector-dependent overall intensity
group/detector-dependent center of scattering function
tunnel splitting
broadening of the tunnel lines
broadening of the elastic line
tunnel circle radius ()
DRAFT: May 29, 2003
Diffusion functions
2-fold jump diffusion
Analytic expression:
1
S(Q, E) = I(Q) A o (Q)(E E c (Q)) + (1 A o (Q) )
(E E c (Q) )2 + 2
where the EISF (elastic-incoherent structure factor) is given by
A o (Q) =
1 + jo (Qd)
2
Model parameters:
I(Q):
:
Ec(Q):
d:
group/detector-dependent overall intensity
half-width at half-maximum of Lorentzian component
group/detector-dependent center of scattering function
jump distance ()
DRAFT: May 29, 2003
3-fold jump diffusion on a circle
Analytic expression:
1
S(Q, E) = I(Q) A o (Q)(E E c (Q)) + (1 A o (Q) )
2
(E E c (Q) ) + 2
where the EISF (elastic-incoherent structure factor) is given by
A o (Q) =
1 + 2 jo (Qr 3 )
3
Model parameters:
I(Q):
:
Ec(Q):
r:
group/detector-dependent overall intensity
half-width at half-maximum of Lorentzian component
group/detector-dependent center of scattering function
jump circle radius ()
DRAFT: May 29, 2003
Translational diffusion
Analytic expression:
S(Q, E) = I(Q)
hDQ 2
1
(E E c (Q) )2 + (hDQ 2 ) 2
Model parameters:
I(Q): group/detector-dependent overall intensity
Ec(Q): group/detector-dependent center of scattering function
D:
diffusion constant (cm2/s)
N-fold jump diffusion on a circle
Analytic expression:
N 1
1
l
S(Q, E) = I(Q) A o (Q)(E E c (Q)) + A l (Q )
2
2
l =1
(E E c (Q) ) + h
l
where the structure factors are given by
A l (Q) =
1 N
2ln
jo (Qrn ) cos
,
N n =1
N
the jump distances are given by
n
rn = 2r sin ,
N
and the inverse correlation times are given by
l
l1 = 2 1 sin 2
N
Model parameters:
I(Q):
Ec(Q):
N:
:
r:
group/detector-dependent overall intensity
group/detector-dependent center of scattering function
number of equivalent sites upon which the particle jumps
inverse correlation time (ns-1 for [E]=eV and ps-1 for [E]=meV)
jump circle radius ()
DRAFT: May 29, 2003
Isotropic Rotational Diffusion
Analytic expression:
A (Q)
S(Q, E) = I(Q) A o (Q)(E E c (Q)) + i
h 2
i =1
2
+ (E E c (Q) )
h
i
where
i1 = i(i + 1)D R , and the structure factors are given by
A o (Q) = jo2 (Qr ) and A i (Q) = (2i + 1) ji2 (Qr ) .
The jl ( x ) are the spherical Bessel functions of the first kind of order l.
Model parameters:
I(Q):
Ec(Q):
DR:
r:
group/detector-dependent overall intensity
group/detector-dependent center of scattering function
rotational diffusion constant (ns-1 for [E] in eV and ps-1 for [E] in meV).
radius of sphere ()
DRAFT: May 29, 2003
Diffusion in a Sphere
(F. Volino and A.J. Dianoux, Molecular Physics 41, 271-279 (1980))
Analytic expression:
(x ln )2 hD 2
0
(2l + 1) l
a
S(Q, E) = I(Q) A 0 (Q)(E E c (Q)) +
A n (Q)
{l , n }{0 , 0}
(x l )2 hD
2
n
+ (E E c (Q) )
a 2
where the structure factors are given by
2
3 j (Qa )
A (Q) = 1
,
Qa
0
0
( )
3 2 l x ln l(l + 1)
A (Q) = jl ( x n )
, Qa = x ln
2
l
2
xn
2
l
n
A ln (Q) =
6(x
(x )
l 2
n
( )
Qajl+1 (Qa ) ljl (Qa )
l
, Qa x n
2
l 2
(Qa ) (x n )
l(l + 1)
l 2
n
the x ln are found as described in the reference above and
the jl ( x ) are the spherical Bessel functions of the first kind of order l.
Model parameters:
I(Q):
Ec(Q):
a:
D:
group/detector-dependent overall intensity
group/detector-dependent center of scattering function
radius of sphere ()
diffusion constant (cm2/s).
DRAFT: May 29, 2003
Bounded Jump Diffusion
(P.L. Hall and D. K. Ross, Molecular Physics 42, 673-682 (1981))
Analytic expression:
A (Q )
n
S(Q, E) = I(Q) A o (Q)(E E c (Q)) + n
2
2
n + (E E c (Q) )
n =1
where
1
2
n = 1 exp (nro / L ) , and the structure factors are given by
2
(2QL) 2
1 (1) n cos(QL) .
A o (Q) = jo2 (QL / 2) and A n (Q) =
2
2 2
(QL) (n)
Model parameters:
I(Q):
Ec(Q):
:
ro :
L:
group/detector-dependent overall intensity
group/detector-dependent center of scattering function
half-width at half-maximum
jump distance ()
confinement distance
DRAFT: May 29, 2003
Miscellaneous functions
Background
Analytic expression:
S(Q, E) = M(Q, E)E + B(Q, E)
Model parameters:
M(Q,E):
B(Q,E):
slope of the background
offset
Gaussian
Analytic expression:
S(Q, E) =
1 E E c (Q )
exp
I(Q)
Model parameters:
I(Q): group/detector-dependent overall intensity
Ec(Q): group/detector-dependent center of scattering function
F(Q): full-width at half-maximum (= 2.354(Q))
Lorentzian
Analytic expression:
S(Q, E) =
I( Q ) ( Q )
1
(E E c (Q)) 2 + (Q) 2
Model parameters:
I(Q): group/detector-dependent overall intensity
Ec(Q): group/detector-dependent center of scattering function
F(Q): full-width at half-maximum (= 2(Q))
Anda mungkin juga menyukai
- ShortCut Buttons - Safe CSIDokumen14 halamanShortCut Buttons - Safe CSIGerry Triaz100% (2)
- SPSS For Beginners: An Illustrative Step-by-Step Approach to Analyzing Statistical dataDari EverandSPSS For Beginners: An Illustrative Step-by-Step Approach to Analyzing Statistical dataBelum ada peringkat
- Finite Element Method Using Pro/Engineer and Ansys: Step 1. Make The PartDokumen7 halamanFinite Element Method Using Pro/Engineer and Ansys: Step 1. Make The PartRithesh Baliga BBelum ada peringkat
- Tableau Training Manual 9.0 Basic Version: This Via Tableau Training Manual Was Created for Both New and IntermediateDari EverandTableau Training Manual 9.0 Basic Version: This Via Tableau Training Manual Was Created for Both New and IntermediatePenilaian: 3 dari 5 bintang3/5 (1)
- Regress It 2020 User ManualDokumen15 halamanRegress It 2020 User ManualIrnanda PratiwiBelum ada peringkat
- Tutorial Surfer 8Dokumen35 halamanTutorial Surfer 8Irwan Tectona RamadhanBelum ada peringkat
- Nemo AnalyserDokumen22 halamanNemo AnalyserNguyễn Hùng SơnBelum ada peringkat
- Etabs TutorialDokumen68 halamanEtabs TutorialBenny Sofyan HartadiBelum ada peringkat
- Tutorial GSASDokumen4 halamanTutorial GSASlina suryantiBelum ada peringkat
- Computer-Controlled Systems: Theory and Design, Third EditionDari EverandComputer-Controlled Systems: Theory and Design, Third EditionPenilaian: 3 dari 5 bintang3/5 (4)
- Emeraude v5.20 Update NotesDokumen15 halamanEmeraude v5.20 Update NotesMuntadher MejthabBelum ada peringkat
- QPA TrainingDokumen13 halamanQPA TrainingsenthilkumarBelum ada peringkat
- Plot XYDokumen8 halamanPlot XYumairnaeem_90Belum ada peringkat
- Dan Dillon Homework Problem #7 - Part 2 STAT 582, Statistical Consulting and Collaboration Dr. JenningsDokumen12 halamanDan Dillon Homework Problem #7 - Part 2 STAT 582, Statistical Consulting and Collaboration Dr. JenningspamfrrBelum ada peringkat
- SPSS HandoutDokumen43 halamanSPSS HandoutÖrdinärySoel100% (1)
- Coda PackDokumen10 halamanCoda PackEdisson Vega100% (1)
- 194 Sample ChapterDokumen27 halaman194 Sample ChapterVikas TiwariBelum ada peringkat
- IBM SPSS Statistics 23 Part 1 - Descriptive StatisticsDokumen18 halamanIBM SPSS Statistics 23 Part 1 - Descriptive StatisticsRitesh SrivastavaBelum ada peringkat
- TC 25 LabEx 1A Intro QGISDokumen8 halamanTC 25 LabEx 1A Intro QGISKenneth T. GuillermoBelum ada peringkat
- Tutorial M MengDokumen197 halamanTutorial M Mengjorgeal958Belum ada peringkat
- Plot XYDokumen10 halamanPlot XYbubo28Belum ada peringkat
- Quick HelpDokumen12 halamanQuick HelpLeo TDBelum ada peringkat
- Data Spikes HiddenDokumen13 halamanData Spikes HiddenhanakistBelum ada peringkat
- TUTORIAL MIDAS GEN 3-D Simple 2-Bay FrameDokumen52 halamanTUTORIAL MIDAS GEN 3-D Simple 2-Bay FrameGiuseppina Sanzone100% (4)
- Eviews PDFDokumen18 halamanEviews PDFaftab20Belum ada peringkat
- Amare - Actions For Marine Protected AreasDokumen15 halamanAmare - Actions For Marine Protected Areasmiftahus siddiqBelum ada peringkat
- KogeoDokumen9 halamanKogeoRamdan YassinBelum ada peringkat
- Mac User Manual - ENGDokumen29 halamanMac User Manual - ENGSalgar DaniloBelum ada peringkat
- ABAQUS Tutorial-Steel PlateDokumen69 halamanABAQUS Tutorial-Steel PlateRabee ShammasBelum ada peringkat
- Finite Element Method Using Pro ENGINEER and ANSYSDokumen11 halamanFinite Element Method Using Pro ENGINEER and ANSYSsunil481Belum ada peringkat
- Fixturlaser Documenter User ManualDokumen58 halamanFixturlaser Documenter User ManualMicroficheBelum ada peringkat
- Introduction To Arcgis: Before You BeginDokumen17 halamanIntroduction To Arcgis: Before You BeginEnoch ArdenBelum ada peringkat
- 3D Shape Rolling ManualDokumen49 halaman3D Shape Rolling ManualWAOLIVEIRA02Belum ada peringkat
- Figure 1. - Credits Screen For BLOCPLAN ForDokumen25 halamanFigure 1. - Credits Screen For BLOCPLAN ForjilliangatheBelum ada peringkat
- Tutorial For Risa Educational: C.M. Uang and K.M. LeetDokumen18 halamanTutorial For Risa Educational: C.M. Uang and K.M. LeetfabianoramiroBelum ada peringkat
- Data Cube: A Relational Aggregation Operator Generalizing Group-By, Cross-Tab, and Sub-TotalsDokumen25 halamanData Cube: A Relational Aggregation Operator Generalizing Group-By, Cross-Tab, and Sub-TotalsJosé A. Ferreira QueimadaBelum ada peringkat
- ATPDraw 5 User Manual UpdatesDokumen51 halamanATPDraw 5 User Manual UpdatesdoniluzBelum ada peringkat
- Experimental WorksheetDokumen8 halamanExperimental WorksheetAlPHA NiNjABelum ada peringkat
- What'S New in Sap Businessobjects Xcelsius 2008 Sp3?: Timo Elliott 31 CommentsDokumen14 halamanWhat'S New in Sap Businessobjects Xcelsius 2008 Sp3?: Timo Elliott 31 CommentsAmanda NandamBelum ada peringkat
- Help AnClimDokumen12 halamanHelp AnClimhcsrecsBelum ada peringkat
- Whats New in FLOW 3D v11.2Dokumen5 halamanWhats New in FLOW 3D v11.2Ahmad HelmiBelum ada peringkat
- Interactive Visualization of Healthcare Data Using TableauDokumen6 halamanInteractive Visualization of Healthcare Data Using TableauJohn JBelum ada peringkat
- An Introduction To Point Pattern Analysis Using CrimestatDokumen19 halamanAn Introduction To Point Pattern Analysis Using CrimestatWelman Rosa AlvaradoBelum ada peringkat
- Hands-On Exercise 2Dokumen11 halamanHands-On Exercise 2Abdulfatai YakubBelum ada peringkat
- Simulase DesignerDokumen9 halamanSimulase Designerdeepalakshmi chandrasekaranBelum ada peringkat
- Lesson 34 - Parameters & RelationsDokumen14 halamanLesson 34 - Parameters & RelationskrongdakBelum ada peringkat
- Aedsys Program: 2002 - Jack D Mattingly, PH.DDokumen34 halamanAedsys Program: 2002 - Jack D Mattingly, PH.Dİlker ÇirkinBelum ada peringkat
- Top 10 Tableau Interview QuestionsDokumen21 halamanTop 10 Tableau Interview QuestionsPata nahiBelum ada peringkat
- Lab Handout - v04 Tableau 01 - Beyond Basic VisualizationDokumen16 halamanLab Handout - v04 Tableau 01 - Beyond Basic VisualizationhammadBelum ada peringkat
- En Tanagra Excel AddIn PDFDokumen9 halamanEn Tanagra Excel AddIn PDFMartin MurciegoBelum ada peringkat
- HP Advanced Design System: Helpful HintsDokumen13 halamanHP Advanced Design System: Helpful HintsyazorcanBelum ada peringkat
- Centurion Data ManagmentDokumen5 halamanCenturion Data ManagmentDIEGO MARTIN MAESOBelum ada peringkat
- Lab Assignment 1: Tutorial For Statgraphics Plus: Ie 355: Quality and Applied Statistics IDokumen10 halamanLab Assignment 1: Tutorial For Statgraphics Plus: Ie 355: Quality and Applied Statistics Ihuynh dungBelum ada peringkat
- Manual SurferDokumen19 halamanManual SurferVincent NeyaBelum ada peringkat
- 9 The Plotting Program Plotxy: Short Description and Operating Instructions (Rel. March 2000)Dokumen8 halaman9 The Plotting Program Plotxy: Short Description and Operating Instructions (Rel. March 2000)Ionutz AxuBelum ada peringkat
- Risk Assessment Using Design Review Based On Failure ModeDokumen6 halamanRisk Assessment Using Design Review Based On Failure ModePaulo LopesBelum ada peringkat
- Fields of Application: About The TestsDokumen47 halamanFields of Application: About The TestsStefan IonelBelum ada peringkat
- WSDOT FOP For AASHTO T 106Dokumen14 halamanWSDOT FOP For AASHTO T 106malaya tripathyBelum ada peringkat
- Laboratory Report - EvaporationDokumen14 halamanLaboratory Report - EvaporationWayne Tandingan0% (1)
- Per g26 Pub 32704 Touchstone AssessmentQPHTMLMode1 32704O236 32704O236S10D1795 17060852160988057 JC1601372310008 32704O236S10D1795E1.html#Dokumen37 halamanPer g26 Pub 32704 Touchstone AssessmentQPHTMLMode1 32704O236 32704O236S10D1795 17060852160988057 JC1601372310008 32704O236S10D1795E1.html#Sandip pawarBelum ada peringkat
- Price Schedule Wapcos Limited Quoting Sheet For The Bidder: Description of Work Unit QuantityDokumen2 halamanPrice Schedule Wapcos Limited Quoting Sheet For The Bidder: Description of Work Unit QuantityBidyut Senapati - WAPCOSBelum ada peringkat
- Drilling and Testing Quality Control Plan - I-35 Project - RevisedDokumen49 halamanDrilling and Testing Quality Control Plan - I-35 Project - RevisedEdwin Antonio Idme Arestegui100% (1)
- Regression AnalysisDokumen25 halamanRegression AnalysisAnkur Sharma100% (1)
- Boiling Point Pure SubstanceDokumen20 halamanBoiling Point Pure SubstanceJay AlbaytarBelum ada peringkat
- Forces and Motion ActivityDokumen5 halamanForces and Motion Activityjanice alquizarBelum ada peringkat
- SR Capital Public SchoolDokumen6 halamanSR Capital Public SchoolLakshya KumarBelum ada peringkat
- Thermal Oil Boiler Vega PDFDokumen2 halamanThermal Oil Boiler Vega PDFrafiradityaBelum ada peringkat
- Scs 210 AmDokumen6 halamanScs 210 AmAntonio CabelloBelum ada peringkat
- Micro Servo RobotDokumen40 halamanMicro Servo Robotlokesh mahor0% (1)
- Filmgate Technical Screen Print InformationDokumen34 halamanFilmgate Technical Screen Print InformationramakrishnafacebookBelum ada peringkat
- A Comparison of Liquid Petroleum Meters For Custody Transfer MeasurementDokumen12 halamanA Comparison of Liquid Petroleum Meters For Custody Transfer MeasurementAmr Guenena100% (2)
- D Angelo Dongre 2009 Practical Use of Multiple Stress Creep and Recovery Test Characterization of Styrene ButadieneDokumen10 halamanD Angelo Dongre 2009 Practical Use of Multiple Stress Creep and Recovery Test Characterization of Styrene Butadienebn23cem3r15Belum ada peringkat
- Topographical Surveys - Direct LevellingDokumen1 halamanTopographical Surveys - Direct LevellingTsegab DereseBelum ada peringkat
- Astm d4945Dokumen7 halamanAstm d4945M.Malyadri ReddyBelum ada peringkat
- SETTLING VELOCITY 2.1 - Calculations of Sedimentation Velocity and Hindered Settling Rate of ParticlesDokumen74 halamanSETTLING VELOCITY 2.1 - Calculations of Sedimentation Velocity and Hindered Settling Rate of ParticlesSonu Singh100% (4)
- Physics: Paper 2Dokumen16 halamanPhysics: Paper 2Mhmd AlrashedBelum ada peringkat
- Installation Manual: Remote User InterfaceDokumen12 halamanInstallation Manual: Remote User InterfaceALEKSANDARBelum ada peringkat
- Soil Structure Interaction Under Dynamic LoadingDokumen9 halamanSoil Structure Interaction Under Dynamic LoadingonurumanBelum ada peringkat
- Permutations and CombinationsDokumen69 halamanPermutations and CombinationsNikhil0% (2)
- At N.p.c.i.l., RawatbhataDokumen30 halamanAt N.p.c.i.l., RawatbhataDevendra SharmaBelum ada peringkat
- Energy Savings and Emissions Reductions For Rewinding and Replacement of Industrial MotorDokumen12 halamanEnergy Savings and Emissions Reductions For Rewinding and Replacement of Industrial Motordedi sanatraBelum ada peringkat
- Delta Industrial Articulated Robot SeriesDokumen15 halamanDelta Industrial Articulated Robot Seriesrobotech automationBelum ada peringkat
- Kids Math - Angles Glossary and TermsDokumen7 halamanKids Math - Angles Glossary and Termssathish11407144Belum ada peringkat
- On Hidden Projection of Plackett Burman Design by Yashi PalDokumen26 halamanOn Hidden Projection of Plackett Burman Design by Yashi PalyashiBelum ada peringkat
- Martini L4 TemperatureControlDokumen11 halamanMartini L4 TemperatureControlJubaer JamiBelum ada peringkat