Final Version - Surf 2016 Rat-Crispr-Prh Poster Se
Diunggah oleh
api-299660206Judul Asli
Hak Cipta
Format Tersedia
Bagikan dokumen Ini
Apakah menurut Anda dokumen ini bermanfaat?
Apakah konten ini tidak pantas?
Laporkan Dokumen IniHak Cipta:
Format Tersedia
Final Version - Surf 2016 Rat-Crispr-Prh Poster Se
Diunggah oleh
api-299660206Hak Cipta:
Format Tersedia
CRISPR/Cas9-Based Development of Novel Transgenic Rat Model of
Congenital Hydrocephalus
A. Scott Emmert1, Crystal Shula1, Francesco Mangano1, Kenneth Campbell1,2, Rolf Stottmann1,3, Yinhuai Chen4, Yueh-Chiang Hu4, and June Goto1
Div. of Pediatric Neurosurgery1, Div. of Developmental Biology2, Div. of Human Genetics3, and Transgenic Animal and Genome Editing Core Facility4, CCHMC
prh
PX459
Results
A
(-) strand sequence
(-) strand sequence
g37
EGP
Figure 1. Electroporation of C6 rat glial cells using the Neon Transfection System.
(+) strand sequence
Figure 5. Design of rat-CRISPR-prh gRNA, primers, and oligonucleotide donor template for introduction of the same single nucleotide
change found in prh mice into rat embryo.
A guide RNA (gRNA) sequence and oligonucleotide donor template
specific to the Ccdc39 gene mutation were designed (Optimized
CRISPR Design) to target a single nucleotide change in the rat
genome. Forward and reverse primers were similarly designed for
insertion of gRNA into a pSpCas9(BB)-2A-Puro (PX459) plasmid
construct.
PX459 plasmid was plated with DH5 competent cells onto LB/amp
agar plates and incubated at 37 for 16 hours. PX459 DNA was
then purified from a transformed bacterial colony and
spectrophotometrically analyzed.
BBSI restriction endonuclease was used to digest PX459, which
was then analyzed for the presence of cleaved bands using gel
electrophoresis. Digested PX459 purified from gel was treated with
alkaline phosphatase.
Diluted insert containing Rat-prh-F and Rat-prh-R primers was
ligated into the PX459 vector to create the PX459-rat-CRISPR-prh
plasmid. DH5 competent cells were transformed with PX459-ratCRISPR-prh, plated, and incubated.
PX459-rat-CRISPR-prh plasmid DNA was purified from the
transformed bacterial culture and analyzed spectrophotometrically.
PX459-rat-CRISPR-prh plasmid was sequenced for correct
insertion and ligation of the gRNA target sequence.
Colonies with positive sequencing results were cultured. PX459-ratCRISPR-prh (rat-prh), PX459, px458-g37 (g37, CRISPR plasmid
positive control), and pEGFP-N1(EGP) DNA was then purified.
Using the Neon Transfection System, C6 cells derived from rat glial
tumor were transfected with rat-prh, PX459, g37, and EGP (Figure
1). EGP fluorescence was used to measure transfection efficiency.
Genomic DNA (gDNA) was extracted from the transfected cells
using the Maxwell 16 System.
Rat-prh and g37 gDNA was amplified with rat-prh assay primers
and g37 assay primers through PCR, and the SURVEYOR
mismatch cleavage assay was used to assess the editing activity of
the rat-CRISPR-prh gRNA construct compared to that of the g37
gRNA positive control (Figure 2).
prh-1
prh-2
g37-2
prh-1
prh-2
g37-1
g37-2
C+G
g37-1
prh-primer
g37-primer
G only
g37-2
prh-primer
prh-1
prh-2
g37-1
g37-2
C+G
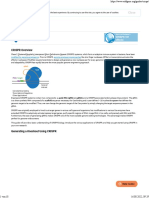
Clustered Regularly Interspaced Short
Palindromic Repeats (CRISPR)
Bacterial immune system modified for
genome engineering
Three components:
1. gRNA
Scaffold: Cas9 binding
Spacer: Genomic target
2. RNA-guided Cas9 endonuclease
3. DNA repair template containing desired
sequence
Cas9-gRNA complex binds genomic target
sequence immediately upstream PAM (5
NGG 3 for S. pyogenes Cas9), Cas9
cleaves target with double stranded break
(DSB), and homology directed repair
Image adapted from Addgene: CRISPR/Cas9 Guide
https://www.addgene.org/CRISPR/guide/
(HDR) generates specific nucleotide
Image adapted from Addgene: CRISPR/Cas9 Guide,
https://www.addgene.org/CRISPR/guide/
change.
Figure 3. CRISPR/Cas9 targeted genome editing using homology directed repair
(HDR).
Progressive hydrocephaly, prh
mouse mutant displaying severe
hydrocephalus at 14-postnatal days
(P14) as compared to wild type
following ENU screen (Stottmann et
al., 2011).
The prh mouse mutants are born with
mild ventriculomegaly but develop
massive hydrocephalus and white
matter degeneration (indicated by
asterisks, F) by P14.
Recently, the possible causal
mutation was found within Ccdc39
Ref. Stottmann et al., Focusing Forward Genetics: A Tripartite ENU Screen for
Neurodevelopmental Mutations in the Mouse Genetics (2011)
gene (unpublished).
Figure 4. Progressive hydrocephaly, prh mouse mutant line isolated in a
forward genetic screen using ENU mutagenesis.
G only
Methods
Background
C only
g37-primer
C only
Figure 2. Process by which SURVEYOR mismatch cleavage assay validates
genetic modification delivered by Cas9-gRNA complex.
prh-1
Image adapted from Surveyor Nuclease Assay, https://en.wikipedia.org/wiki/Surveyor_nuclease_assay
A. Using Optimized CRISPR Design (Zhang Lab, MIT 2015), a guide RNA was designed to target the critical thymine nucleotide required for proper splicing
of the rat Ccdc39 gene (chr2: 139960533A, rn5), which is mutated in the prh mouse model of congenital hydrocephalus. The gRNA sequence must lie
immediately upstream the PAM sequence (5 TGG 3) for Cas9 to bind target DNA. Guide #4 was selected as the most efficient gRNA design for its high ontarget quality score and low off-target activity score. Rat-CRISPR-prh forward and reverse primers were similarly designed to include the Guide #4 gRNA
targeting sequence for insertion using the PX459 vector. B. Negative (PAM-containing) and positive (non-PAM-containing) strand sequences of donor
oligonucleotide repair template were constructed with substitution of thymine to adenine in target sequence. Donor oligonucleotide was designed with
HindIII restriction site as SNP genotyping strategy.
prh-2
Congenital hydrocephalus is a devastating birth defect defined by
the abnormal accumulation of cerebrospinal fluid (CSF) in the brain.
Buildup of CSF places enormous pressure on the infants brain,
ultimately leading to brain damage and mental and physical
problems if left untreated. Although surgical intervention remains the
primary form of treatment for congenital hydrocephalus, it is not a
cure for this condition.
Emergence of the novel CRISPR/Cas9 genome-editing platform
provides an accessible and efficient method for investigating
possible gene mutations responsible for congenital hydrocephalus.
Through whole-genome sequencing, our group recently identified a
splice mutation within Ccdc39 (Coiled-coil domain containing 39)
gene as a strong mutation candidate for the mutant phenotype in a
congenital hydrocephalus mouse model, progressive hydrocephaly,
prh.
Defects in Ccdc39 gene disrupt normal motility of cilia and are the
cause of primary ciliary dyskinesia in humans and dogs.
Using the CRISPR/Cas9 system of targeted genome editing, this
study aims to introduce the same mutation evidenced in the prh
mouse mutant line into the rat genome.
Creation of a prh rat model of congenital hydrocephalus provides a
method to evaluate whether the mutation in Ccdc39 is responsible
for the hydrocephalus phenotype. Also, the prh rat model presents
opportunities to explore new surgical and imaging techniques and
to analyze the molecular functions of the Ccdc39 gene in rodent
brain development.
Methods (cont.)
g37-1
Introduction
900
800
700
900
800
700
600
500
400
600
500
300
400
200
300
100
200
100
2% gel 30 min
2% gel 1 hr
Figure 6. SURVEYOR mismatch cleavage assay confirms ability of PX459-rat-CRISPR-prh plasmid construct to modify genome of C6
rat glial cells following CRISPR/Cas9 genome editing.
Gel electrophoresis analysis of rat-CRISPR-prh and g37 digested DNA products using SURVEYOR mismatch cleavage assay. Prh and g37 DNA amplified
with corresponding g37-T7E1 and prh-T7E1 assay primers display cleaved bands indicative of SNP or indel in region of interest. Prh/prh-primer PCR
product at 888 bp is cleaved into 569 bp and 319 bp bands (red ovals) as initially designed into rat-CRISPR-prh T7E1 assay primers, demonstrating the
introduction of the single nucleotide change from thymine to adenine into rat genome as originally found in prh mice.
Conclusions
PX459-rat-CRISPR-prh plasmid and gRNA construct, which were designed to introduce the point mutation from thymine to adenine found in
prh mice into rat embryos, are capable of modifying the rat genome.
The successful result of this gRNA editing activity test in C6 cells demonstrates that the CRISPR/Cas9 platform can be used to generate a prh
mutant rat model of congenital hydrocephalus. This platform for targeted genome editing provides an efficient and adaptable method for
studying the molecular mechanisms of genetic diseases.
gRNA design programs like Optimized CRISPR Design can be used to create CRISPR/Cas9 experiments that readily and precisely target
nearly any sequence in the genome.
Future Directions
Prh mutant rat line is currently being generated with help from the Transgenic Animal and Genome Editing Core Facility. Zygote injections of
rat-CRISPR-prh plasmid construct, gRNA, and donor oligonucleotide into Sprague Dawley rats are underway.
Rat mutants displaying progressive hydrocephaly, prh phenotype will be genotyped, surgically shunted by neurosurgeons from the Division
of Pediatric Neurosurgery, and analyzed using diffusion tensor imaging (DTI).
Evidence of hydrocephalic prh rats following introduction of the same single nucleotide change found in prh mice would strengthen the
hypothesis of this point mutation as the mutation candidate of the Ccdc39 splice mutation.
Anda mungkin juga menyukai
- Shoe Dog: A Memoir by the Creator of NikeDari EverandShoe Dog: A Memoir by the Creator of NikePenilaian: 4.5 dari 5 bintang4.5/5 (537)
- The Yellow House: A Memoir (2019 National Book Award Winner)Dari EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Penilaian: 4 dari 5 bintang4/5 (98)
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeDari EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifePenilaian: 4 dari 5 bintang4/5 (5794)
- The Little Book of Hygge: Danish Secrets to Happy LivingDari EverandThe Little Book of Hygge: Danish Secrets to Happy LivingPenilaian: 3.5 dari 5 bintang3.5/5 (400)
- Grit: The Power of Passion and PerseveranceDari EverandGrit: The Power of Passion and PerseverancePenilaian: 4 dari 5 bintang4/5 (588)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureDari EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FuturePenilaian: 4.5 dari 5 bintang4.5/5 (474)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryDari EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryPenilaian: 3.5 dari 5 bintang3.5/5 (231)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceDari EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RacePenilaian: 4 dari 5 bintang4/5 (895)
- Team of Rivals: The Political Genius of Abraham LincolnDari EverandTeam of Rivals: The Political Genius of Abraham LincolnPenilaian: 4.5 dari 5 bintang4.5/5 (234)
- Never Split the Difference: Negotiating As If Your Life Depended On ItDari EverandNever Split the Difference: Negotiating As If Your Life Depended On ItPenilaian: 4.5 dari 5 bintang4.5/5 (838)
- The Emperor of All Maladies: A Biography of CancerDari EverandThe Emperor of All Maladies: A Biography of CancerPenilaian: 4.5 dari 5 bintang4.5/5 (271)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaDari EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaPenilaian: 4.5 dari 5 bintang4.5/5 (266)
- On Fire: The (Burning) Case for a Green New DealDari EverandOn Fire: The (Burning) Case for a Green New DealPenilaian: 4 dari 5 bintang4/5 (74)
- The Unwinding: An Inner History of the New AmericaDari EverandThe Unwinding: An Inner History of the New AmericaPenilaian: 4 dari 5 bintang4/5 (45)
- Rise of ISIS: A Threat We Can't IgnoreDari EverandRise of ISIS: A Threat We Can't IgnorePenilaian: 3.5 dari 5 bintang3.5/5 (137)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersDari EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersPenilaian: 4.5 dari 5 bintang4.5/5 (345)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyDari EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyPenilaian: 3.5 dari 5 bintang3.5/5 (2259)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreDari EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You ArePenilaian: 4 dari 5 bintang4/5 (1090)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)Dari EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Penilaian: 4.5 dari 5 bintang4.5/5 (121)
- Her Body and Other Parties: StoriesDari EverandHer Body and Other Parties: StoriesPenilaian: 4 dari 5 bintang4/5 (821)
- Rewriting A GenomeDokumen2 halamanRewriting A GenomeUjwalBelum ada peringkat
- The Gene - Siddhartha MukherjeeDokumen38 halamanThe Gene - Siddhartha MukherjeejohndevonrileyBelum ada peringkat
- Springer Protocols Handbooks M Tofazzal Islam Pankaj K Bhowmik Kutubuddin A Molla - CRISPR-Cas Methods-Springer US 20Dokumen288 halamanSpringer Protocols Handbooks M Tofazzal Islam Pankaj K Bhowmik Kutubuddin A Molla - CRISPR-Cas Methods-Springer US 20Никита Ваулин100% (1)
- Crispr Cas Ethics ArticleDokumen10 halamanCrispr Cas Ethics ArticleJuan Carlos RamírezBelum ada peringkat
- CRISPR - YarrowiaDokumen9 halamanCRISPR - YarrowiaRegi RGBelum ada peringkat
- Addgene CRISPR GuideDokumen18 halamanAddgene CRISPR GuideThomasBelum ada peringkat
- The CRISPR SystemDokumen13 halamanThe CRISPR SystemNádia GavilánBelum ada peringkat
- Science - and - Technology PT 365Dokumen83 halamanScience - and - Technology PT 365mobile legend gaming farhan the fab.Belum ada peringkat
- Barrangou Et Al. - 2007 - CRISPR Provides Acquired Resistance Against Viruses in Prokaryotes PDFDokumen5 halamanBarrangou Et Al. - 2007 - CRISPR Provides Acquired Resistance Against Viruses in Prokaryotes PDFflashjetBelum ada peringkat
- The Future of Gene EditingDokumen23 halamanThe Future of Gene EditingAnurag KunwarBelum ada peringkat
- Content ServerDokumen12 halamanContent ServerMARIA DANIELA PADILLA MUÑOZBelum ada peringkat
- Crispr AssignmentDokumen7 halamanCrispr AssignmentJuan RamirezBelum ada peringkat
- CRISPR Gene Editing PDFDokumen356 halamanCRISPR Gene Editing PDFCoimbra Rojas100% (5)
- (Debate) Sex EducationDokumen5 halaman(Debate) Sex EducationHeath Bag PkyBelum ada peringkat
- MGI Bio Revolution Report May 2020Dokumen200 halamanMGI Bio Revolution Report May 2020Syed Arshad JamalBelum ada peringkat
- Kadhem Fatima 218809459 Assgn3Dokumen5 halamanKadhem Fatima 218809459 Assgn3api-663622821Belum ada peringkat
- Lentilactobacillus Buchneri Domination During TheDokumen7 halamanLentilactobacillus Buchneri Domination During TheSamuel Ayeh OseiBelum ada peringkat
- Review: The Biology of CRISPR-Cas: Backward and ForwardDokumen21 halamanReview: The Biology of CRISPR-Cas: Backward and ForwardNiharika ChaudharyBelum ada peringkat
- Applications of Biotechnology in HealthDokumen21 halamanApplications of Biotechnology in Healtherinliannamanalad4210Belum ada peringkat
- A CRISPR Way To Block PERVs - Engineering OrgansDokumen3 halamanA CRISPR Way To Block PERVs - Engineering OrgansFrancisco Baca DejoBelum ada peringkat
- ML Full Practice Test 5Dokumen20 halamanML Full Practice Test 5dilyahanim707100% (1)
- Crispr2019 PDFDokumen24 halamanCrispr2019 PDFsumedhaBelum ada peringkat
- VisionIAS PT 365 December 2024 Science and TechnologyDokumen112 halamanVisionIAS PT 365 December 2024 Science and TechnologySourabh SinghBelum ada peringkat
- Synthetic Guide RNA For CRISPR Genome EditingDokumen9 halamanSynthetic Guide RNA For CRISPR Genome EditinggiacummoBelum ada peringkat
- Genome EditingDokumen12 halamanGenome EditingDahlia StudioBelum ada peringkat
- Nihms 1544880Dokumen51 halamanNihms 1544880Robert StryjakBelum ada peringkat
- Crispr Cas A Laboratory ManualDokumen10 halamanCrispr Cas A Laboratory Manualʜᴏsᴅᴇ ¿Belum ada peringkat
- Earth and Life Science Genetic EngineeringDokumen66 halamanEarth and Life Science Genetic EngineeringJisho 恐怖 Lomotos75% (4)
- Reflective Essay: Writing in The Genetics DiscourseDokumen5 halamanReflective Essay: Writing in The Genetics DiscourseAnonymous AY6XDZHBxPBelum ada peringkat
- Manuscript Introduction - Plain TextDokumen2 halamanManuscript Introduction - Plain Textد. محمد قاسمBelum ada peringkat